Characterizing the Skin Microbiome, One Genome at a Time
Julie Segre Delivers the Anita Roberts Lecture
We often associate microbes with disease. But the Human Microbiome Project has shown that people live in a mutually beneficial relationship with trillions of microbes on and in their bodies. Human diseases affecting the mouth (cavities) and gastrointestinal tract (inflammatory bowel disease) as well as more complex diseases such as premature birth or type 2 diabetes are associated with alterations in the microbiome. And now, clinical studies are investigating whether those modifications might even drive some features of disease.
Julie Segre
Senior Investigator Julie A. Segre (National Human Genome Research Institute, NHGRI), who is exploring the skin microbiome, explained the dichotomy in her Anita Roberts Lecture titled “Human Microbiome: Friend or Foe” on November 3, 2020. She uses genome sequencing and computational methods to characterize the healthy skin microbiome to better understand the pathology of and design novel therapies for skin-related disorders such as atopic dermatitis (AD), or eczema.
So far there is no cure for AD although symptoms can be treated with topical application of emollients, corticosteroids, and antimicrobials. Segre’s lab, however, is investigating what causes it. She discovered that, in AD, there is a decrease in the skin’s microbial diversity and an increase in two species of the Staphylococcus bacteria—S. aureus and S. epidermidis. In collaboration with her long-time clinical partner, Heidi Kong (National Institute of Arthritis and Musculoskeletal and Skin Diseases), Segre’s lab found that topically applying Staphylococcal strains from patients with severe flare-ups of AD cause skin and immune changes in mice; strains from patients with milder disease cause no symptoms in mice. Segre envisions developing patient-tailored treatments by understanding the basic mechanisms of host-microbial interactions in mice first.
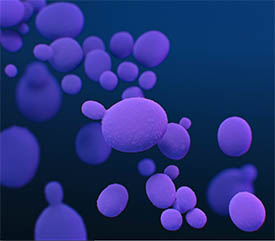
CREDIT: MEDICAL ILLUSTRATOR: STEPHANIE ROSSOW, CDC
Julie Segre is studying Candida auris, a fungus that is emerging as a global health threat because it is multi-drug resistant and has caused outbreaks in nursing homes, according to the Centers for Disease Control and Prevention (CDC). Shown: A medical illustration of C. auris fungal organisms, presented in the CDC publication entitled, Antibiotic Resistance Threats in the United States, 2019.
The skin microbiome also includes a diversity of fungi, which Segre is also studying. In particular, she is focusing on Candida auris, a fungus that is emerging as a global health threat because it is multi-drug resistant and has caused outbreaks in nursing homes, according to the Centers for Disease Control and Prevention. First seen in patients in 2008 in Asia, Africa, and South America, C. auris has spread throughout the world. The fungus can colonize skin for months to years; it can easily spread; and it also causes difficult–to–treat bloodstream infections. In a recent study, Segre’s team explored the spread of C. auris in a nursing facility to identify sites of colonization and changes in the skin microbiome associated with colonization. She hopes that these insights will help to halt the spread of this emerging human fungal pathogen. (Open Forum Infect Dis 6(Suppl 2):S25–S26, 2019)
She counsels young investigators and researchers to develop quantitative skills and build collaborations, advice she has followed herself. Among her quantitative skills is the ability to use computational tools effectively. “I was not afraid to take ownership of analyzing the humungous amount of data that sequencing experiments generate,” she said. And she has had many successful long-term collaborations: with Kong; Yasmine Belkaid (National Institute of Allergy and Infectious Diseases, NIAID), who developed the mouse models that Segre used and is trying to understand the mechanisms controlling host–microbe interactions on the skin and in the gut; Helen Su (NIAID), who is evaluating the microbial colonization of patients with primary immune deficiency; and many others.
Segre graduated summa cum laude with a B.S. in mathematics from Amherst College (Amherst, Massachusetts) in 1987 and earned a Ph.D. in biology from the Massachusetts Institute of Technology (MIT; Cambridge, Massachusetts) in 1995. She was the first Ph.D. student of genome-sequencing pioneer Eric Lander at MIT and did her postdoctoral training with cell biologist Elaine Fuchs at the University of Chicago. Segre joined NHGRI as a senior investigator in 2000 and was tenured in 2007. She now leads NHGRI’s Translational and Functional Genomics Branch and the Microbial Genomics Section.
Like Anita Roberts, Segre is an exceptional scientist, a great leader and mentor, and a kind and considerate human being. In addition to performing cutting-edge science, Segre is committed to nurturing a more inclusive and diverse workforce. She is a founding member of the NIH Anti-Harassment Committee and the NIH Equity Committee. She also acknowledges the support of the Women Scientists Advisors in advocating for women scientists at NIH.
The “Anita B. Roberts Lecture Series: Distinguished Women Scientists at NIH” honors her research contributions. Roberts, who spent 30 years at NCI before her untimely death in 2006 from gastric cancer, was known for her groundbreaking work on transforming growth factor–beta. To watch a videocast of Segre’s lecture “Human Microbiome: Friend or Foe,” held on November 3, 2020, go to https://videocast.nih.gov/watch=37780.

Sunita Chopra is a visiting postdoctoral fellow in the National Cancer Institute’s Radiation Oncology Branch, where she studies radiation-responsive coding and noncoding RNA signatures in the blood of whole-body-irradiated animal models. She plans to transition into a writing-intensive career after finishing her training at NIH. Outside of work, she enjoys being outdoors, cooking, listening to Bollywood songs, and being with friends.
This page was last updated on Thursday, March 10, 2022
