Research Briefs
Read about NIH scientific advances and discoveries by intramural scientists:
- NEI: RESEARCH REVEALS NEW ASPECTS OF VISUAL PROCESSING
- NINDS, NCATS: COMMON ANTIBIOTIC SHOWS PROMISE AGAINST ZIKA INFECTIONS
- NIA, NINDS: TWO NERVOUS SYSTEM DISORDERS LINKED TO MUTATION ASSOCIATED WITH HUNTINGTON DISEASE
- NCCIH, NINDS: PIEZO2 GENE HELPS US KNOW WHEN TO URINATE
- NIEHS: DOUBLE-STRAND DNA BREAKS REPAIRED BY POLYMERASE MU
- NIEHS, NCI: BREAST CANCER, AIR POLLUTION LINKED BY NEW MARKER
- NCI: STUDY OF “EXCEPTIONAL RESPONDERS” YIELDS CLUES TO CANCER AND POTENTIAL TREATMENT
- NIAID RML: EXPERIMENTAL VACCINE PROTECTS AGAINST DEADLY TICKBORNE VIRUS
- NINDS: THE GUT TRAINS THE IMMUNE SYSTEM TO PROTECT THE BRAIN
- NICHD: GENE IN MICE THAT CONTROLS FOOD CRAVINGS AND DESIRE TO EXERCISE
NEI: RESEARCH REVEALS NEW ASPECTS OF VISUAL PROCESSING
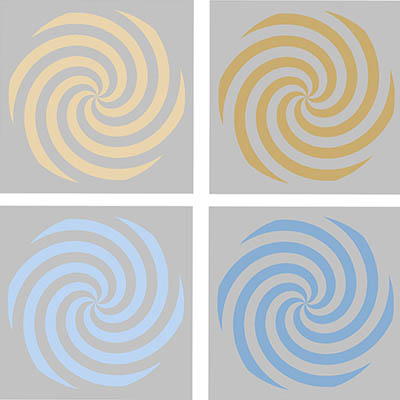

CREDIT: BEVIL CONWAY, NEI
In an NEI study, magnetoencephalography recordings revealed that study participants showed unique patterns of brain activity for each color they viewed. Shown: Colored stimuli in yellow (top) and blue (bottom). Light luminance level versions are on the left; dark versions on the right. Volunteers used a variety of names for the upper stimuli, such as “yellow” for the left and “brown” for the right, but consistently used “blue” for both the lower stimuli.
Color perception, the process by which the brain receives information from cones in the retina and converts the signals into what we perceive as color vision, is largely a mystery. NEI researchers have decoded brain maps of human color perception and learned how color processing is organized in the brain. The scientists used a technique called magnetoencephalography (MEG) that involves placing an array of sensors around the head to noninvasively record the tiny magnetic fields that accompany brain activity. As study participants viewed specially designed color images and reported the colors they saw, the recordings revealed unique patterns of brain activity for each color. The researchers were even able to predict from MEG recordings what color a participant was viewing.
In addition, the study revealed correlations between neurological signals and patterns in color naming. For example, the researchers found greater variation in brain activity between light and dark hues of warm colors (yellows, reds, oranges, browns) than for the same hues of cool colors (blues, greens), which supports the empirical observation that different hues of warm colors are more precisely named than cool colors. The findings indicate that the color-naming conventions consistent across many languages and cultures may have a fundamental neurological basis. These results reveal new insights into the physiological basis for color and how brain activity measurements can be used to model color perception. The study may also have implications for the development of machine-brain interfaces for visual prosthetics. (NIH authors: I.A. Rosenthal, S.R. Singh, K.L. Hermann, and B.R. Conway, Curr Biol 31:1–12, 2020; DOI:10.1016/j.cub.2020.10.062)
NINDS, NCATS: COMMON ANTIBIOTIC SHOWS PROMISE AGAINST ZIKA INFECTIONS
CREDIT: AVI NATH LAB, NINDS
NIH scientists found that the commonly used antibiotic methacycline may be effective at combating the neurological problems caused by Zika virus infections. This is a picture of a Zika-infected mouse brain from the study.
The global outbreak of Zika virus, primarily spread by the Aedes aegypti mosquito, emerged as a public health emergency in 2015 and 2016 with at least 60 countries reporting active cases. When infection occurs during pregnancy, the virus may cross the placental barrier and cause babies to be born with microcephaly (abnormally small heads). Some adults develop severe neurological disorders such as Guillain-Barré syndrome, encephalitis, and myelitis. Scientists in NINDS, in NCATS, and at Georgetown University Medical Center (Washington, D.C.) collaborated on the study and used a high-throughput technique to screen more than 130,000 compounds to identify several inhibitors of Zika virus infection.
The researchers looked for drugs that block the activity of proteins called NS2B-NS3 Zika virus protease and prevent the virus from reproducing. Study leader Rachel Abrams (NINDS), in Avindra Nath’s lab, worked with other scientists and found that the widely available and commonly used tetracycline-based drugs, which includes the antibiotic methacycline, may be effective at blocking Zika virus infection in mice and reduced some of the resulting neurodevelopmental problems. Tetracyclines are FDA-approved drugs that are known to cross the placenta of pregnant women. According to Abrams, because tetracycline-based antibiotics are widely used, their potential can be rapidly tested in clinical trials. (NIH authors: R.P.M. Abrams, A. Yasgar, M.-H. Lee, D. Dorjsuren, R. T. Eastman, N. Malik, A.V. Zakharov, W. Li, M. Bachani, K. Brimacombe, J.P. Steiner, M.D. Hall, A. Jadhav, A. Simeonov, and A. Nath, Proc Natl Acad Sci USA 117:31365–31375, 2020; DOI:10.1073/pnas.2005463117)
NIA, NINDS: TWO NERVOUS SYSTEM DISORDERS LINKED TO MUTATION ASSOCIATED WITH HUNTINGTON DISEASE
NIA and NINDS researchers led an international study that discovered a surprising connection between two nervous-system disorders—frontotemporal dementia (FTD) and amyotrophic lateral sclerosis (ALS)—and a mutation in the huntingtin gene (HTT), which causes Huntington disease.
By comparing the DNA of ALS and FTD patients with DNA from aged-matched healthy individuals, the scientists found repeat expansions (abnormal repetition of certain DNA building blocks) in the HTT gene in a small subset of ALS and FTD patients, who did not show the classic symptoms of Huntington disease.
“Our patients simply don’t match a textbook definition of disease when it comes to which mutation produces which symptoms,” said Sonja Scholz (NINDS), one of the investigators of the study. “Here we have patients carrying a pathogenic huntingtin mutation but who present with FTD or ALS symptoms.”
The authors believe that adding genetic screening for the mutation could increase diagnostic accuracy for patients showing symptoms of ALS or FTD and make personalized therapies possible for the small subset of patients who have the mutation. Advanced clinical trials targeting the mutation are already underway. (NIH authors: R. Dewan, R. Chia, J. Ding, Y. Abramzon, S. Ahmed, M.S. Sabir, M.K. Portley, A.B. Singleton, T. Tanaka, L. Ferrucci, S.M. Resnick, J.R. Gibbs, S.W. Scholz, and B.J. Traynor, Neuron 109:1–13, 2021; DOI:https://doi.org/10.1016/j.neuron.2020.11.005)
NCCIH, NINDS: PIEZO2 GENE HELPS US KNOW WHEN TO URINATE
CREDIT: ARDEM PATAPOUTIAN LAB, SCRIPPS RESEARCH INSTITUTE
Researchers discovered that the PIEZO2 gene may help us sense when our bladders are full and it is time to urinate. Shown: Immunofluorescence microscopy image of a mouse bladder (tissue stained red; nuclei stained blue).
Scientists at NIH and the Scripps Research Institute (La Jolla, California) discovered that a gene, called PIEZO2, may be responsible for the powerful urge to urinate that we normally feel several times a day. The results suggest that the gene helps at least two different types of cells sense when our bladders are full and need to be emptied. The study involved both mice and patients who are part of a clinical trial at the NIH Clinical Center.
The PIEZO2 and PIEZO1 genes were discovered in 2010 by NIH-funded investigators at Scripps. Since then, NIH teams led by Alex Chesler (NCCIH) and Carsten Bönnemann (NINDS) have shown that, in humans, the PIEZO2 gene may play many roles including controlling the sense of touch, vibration, and proprioception (the conscious as well as unconscious awareness of one’s body in space). In 2015, the NIH researchers determined that people who were born with disabling mutations in their PIEZO2 had no sense of proprioception and couldn’t feel some forms of touch and pain.
In the new study—the first in-depth examination of human subjects with PIEZO2 defects—the NIH investigators discovered that people without PIEZO2 function also have impaired bladder control and problems with urination. To understand the mechanics of PIEZO2’s role, the researchers examined mice. Using a combination of histological techniques and an advanced real-time in vivo imaging platform developed at NIH, the investigators found that the PIEZO2 gene was largely expressed in bladder-innervating neurons and special “umbrella cells” that line the bladder lumen. Compared with normal mice, the PIEZO2—deficient mice were less sensitive to bladder filling and more volume was needed to initiate bladder contractions; showed irregular timing of urination; and had significantly more pressure on urinary bladder muscles during micturition (the act of urinating). Collectively, these results demonstrate that PIEZO2 sets the bladder stretch sensitivity in the lower urinary tract to achieve micturition in a timely manner. Researchers will continue their work and explore the clinical implications for the millions suffering from urinary-control problems. (NIH authors: D. Saade, N. Ghitani, M. Szczot, T. Ogata, C.G. Bönneman, and A.T. Chesler, Nature 588:290–295, 2020; DOI:10.1038/s41586-020-2830-7)
NIEHS: DOUBLE-STRAND DNA BREAKS REPAIRED BY POLYMERASE MU
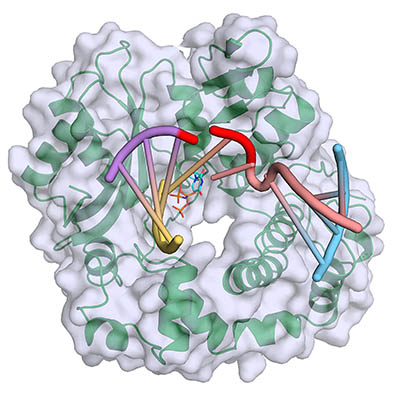
CREDIT: NIEHS
Polymerase mu (green, with gray surface) binds and bridges a DNA double-strand break, filling gaps at the break site (red) with incoming complementary nucleotides (blue). Yellow and purple strands represent the upstream DNA duplex, and pink and blue strands represent the downstream DNA duplex.
NIEHS researchers have taken the first snapshot of polymerase mu (Pol-mu) as it bridges a double-strand break (DSB), the most toxic form of DNA damage. Persistently unrepaired DSBs can lead to cancers and other diseases. The findings may aid the understanding of cancer and cancer therapeutics because cancer cells depend heavily on this type of repair. Pol-mu, one of the few enzymes that helps repair these breaks, is capable of handling DSBs that have jagged, unpaired ends. These snapshots allowed the researchers to gain critical insight into how the enzyme makes repair of DSBs possible. Pol-mu formed a remarkably rigid structure that bridged the two severed strands of DNA. Going forward, the researchers plan to find out how all the enzymes involved in DNA double-strand-break repair work together. (NIH authors: A.M. Kaminski, T.A. Kunkel, L.C. Pedersen, and K. Bebenek, Nat Commun 11:4784, 2020; DOI:10.1038/s41467-020-18506-5) [https://www.nature.com/articles/s41467-020-18506-5]
NIEHS, NCI: BREAST CANCER, AIR POLLUTION LINKED BY NEW MARKER

CREDIT: SHUTTERSTOCK.COM
In addition to PM-2.5, gaseous pollutants were associated with elevated TDLU count, but not in a linear fashion. Shown here, steel mills in Indiana, the state where many of the tissue donors lived. Shown: Steel Mills in Indiana.
Researchers at NIEHS, NCI, and colleagues reported a previously unknown connection between outdoor air pollution and terminal duct lobular units (TDLUs; epithelial structures within the breast that produce milk during lactation), where most breast cancers originate. As women age, their TDLUs naturally shrink and become fewer, a process called involution. Reduced involution is linked to an increased risk of breast cancer. The researchers analyzed more than 1,900 samples of healthy breast tissue donated by women to the Komen Tissue Bank and compared the number and size of TDLUs with air-pollution exposure levels where the women resided. Living in areas with high levels of fine particulate matter (less than 2.5 micrometers in diameter, or PM-2.5), held the strongest association with reduced involution. These findings strengthen the biologic rationale for the air pollution–breast cancer connection. Past studies suggested that TDLU involution may occur at least in part through a hormone-related pathway. The scientists suggested that PM-2.5 may influence changes in breast tissue by disrupting hormone balances. (NIH authors: N.M. Niehoff, A.P. Keil, R.R. Jones, S. Fan, G.L. Gierach, and A.J. White, Breast Cancer Res 22:100, 2020; DOI:10.1186/s13058-020-01339-x)
[BY ERNIE HOOD, NIEHS]
NCI: STUDY OF “EXCEPTIONAL RESPONDERS” YIELDS CLUES TO CANCER AND POTENTIAL TREATMENT

CREDIT: NHGRI
NCI: A genomic study has uncovered molecular changes in patient tumors that may give rise to dramatic and long-lasting responses to cancer therapy.
A small percentage of cancer patients are classified as “exceptional responders” because although they have advanced disease, they survive significantly longer than similar patients. NCI researchers, in collaboration with investigators from other institutions, did a comprehensive analysis of exceptional responders and found molecular changes in the tumors that may provide some explanation.
The study involved 111 exceptional responders (most had difficult-to-treat metastatic cancers) who had received chemotherapy. To analyze the tumor samples, researchers used multiple genomic techniques—to profile DNA mutations, RNA expression levels, DNA methylation, and DNA copy-number changes—and also examined the immune cells in the tumor microenvironment. For nearly a fourth of the patients, the scientists identified broad categories of mechanisms behind the efficient response to chemotherapy treatment: the immune system’s response to tumors; the ability to repair DNA damage; and synthetic lethality (when a combination of deficiencies in two or more genes leads to the death of tumor cells during treatment). In some patients there was an increase in the amount of B lymphocytes that indicated that the body’s immune system was trying to control tumor growth. Two patients had mutations in the DNA-repairing genes BRCA1 or BRCA2; the mutations meant that tumors couldn’t fix damaged DNA and therefore were more susceptible to platinum-based chemotherapy treatments. The researchers noted that more research and additional analytical approaches are needed to describe the molecular underpinnings of the unsolved cases of unexceptional responders. (NCI authors: N. Takebe, Jean C. Zenklusen, R. Tarnuzzer, L.M. McShane, J.V. Tricoli, P.M. Williams, I. Lubensky, G. O’Sullivan-Coyne, E.C. Kohn, R.F. Little, J. White, S. Malik, L. Harris, C. Weil, A.P. Chen, C. Karlovich, B. Rodgers, L. Shankar, P. Jacobs, E.F. Edmondson, J.H. Doroshow, B.A. Conley, S.P. Ivy, and L.M. Staudt, Cancer Cell 39:1–16, 2021; DOI:10.1016/j.ccell.2020.10.015)
NIAID RML: EXPERIMENTAL VACCINE PROTECTS AGAINST DEADLY TICKBORNE VIRUS
NIAID researchers at the Rocky Mountain Labs (Hamilton, Montana) helped colleagues in Europe develop an experimental DNA-based vaccine that is protective against Crimean-Congo hemorrhagic fever virus (CCHFV) in Cynomolgus macaques. The virus, which causes Crimean-Congo hemorrhagic fever, is primarily transmitted by Hyalomma ticks (which are endemic in Eastern Europe, Africa, the Middle East, and parts of Asia) and also through the handling of infected livestock or care of infected patients. According to the World Health Organization, CCGFV infects up to 15,000 people a year; 1 in 8 of those infected develop severe disease with hemorrhagic manifestations; and CCGFV kills about 500 people annually. Global climate change may contribute to the expansion of these ticks’ range leading to the introduction of CCHFV in new regions. A vaccine is needed to protect at-risk populations including people in rural areas, farmers, slaughterhouse workers, and health-care workers.
NIAID researchers tested the experimental DNA-based vaccine, consisting of plasmid expressing the nucleoprotein and viral glycoproteins of CCHFV, on six primates. Each macaque received three inoculations, followed by electroporation (electrical pulses to facilitate the delivery of plasmid DNA and increase immune responses) at three-week intervals. The researchers confirmed through regular blood tests that the vaccine generated protective antibodies against the virus. Then the vaccinated animals were infected with CCHFV and monitored for six days; they did not get sick. Six control animals infected with the virus but not given the vaccine did develop signs of the disease. These results suggest that this vaccine can be advanced to human clinical trials. (NIH authors: D.W. Hawman, K. Meade-White, P.W. Hanley, D. Scott, A. Okumura, and H. Feldmann, Nat Microbiol 2020; DOI:10.1038/s41564-020-00815-6)
NINDS: THE GUT TRAINS THE IMMUNE SYSTEM TO PROTECT THE BRAIN
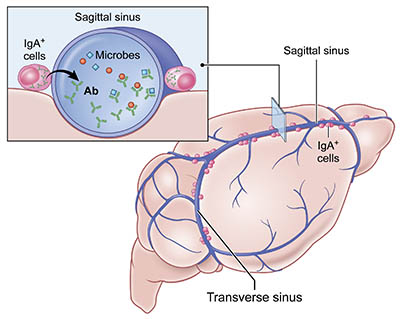
CREDIT: DORIAN MCGAVERN’S LAB, NINDS
NINDS: Gut-educated immune cells protect the brain: Immunoglobulin A (IgA) cells reside in the dura mater along venous sinuses that drain blood from the brain. These cells produce antibodies (Ab) against microbes they first encountered in the gut.
Historically, the brain has been viewed as an immune-privileged site, meaning that it can tolerate foreign antigens without launching an immune response. New research, conducted by researchers at NINDS and Cambridge University (Cambridge, England), has shown that the meninges (a three-membrane barrier that protects the brain and spinal cord from pathogens) contain a diverse population of immune cells, some of which are trained to fight infections by first spending time in the gut.
Unexpectedly, the investigators found that large venous sinuses residing in the meninges of healthy mice contained many antibody-producing plasma cells called immunoglobulin A (IgA) cells. These IgA cells are common on barrier surfaces such as the gut or skin, which are constantly secreting antibodies to protect against the microbes of the outside world. It was surprising, however, to find these IgA cells inhabiting the walls of meningeal sinuses responsible for draining blood from the brain. Observing them in this location suggests that they help protect the brain from foreign intruders that enter the blood.
Compared with the controls, germ-free mice (which do not have their own microbiome) had almost no IgA cells in their meninges. When the researchers reconstituted the gut microbiome in these mice, the population of meningeal IgA cells was restored. This finding suggests that the gut bacteria are important in shaping the antibodies produced by the meningeal IgA cells.
“Because these cells are educated in the gut, anything that influences the gut environment is going to potentially change their number, distribution, and antibody specificity,” said Dorian McGavern, senior investigator and co-author of the paper that appeared in Nature. When treated with oral antibiotics, the population of meningeal IgA cells decreased substantially in the mice. “It shows you that you can give antibiotics to treat one microbe, but that opens up susceptibility in your immunological defense in another place you didn’t consider,” he said.
Researchers compared the DNA sequences of IgA cell antibodies in the meninges with those from the intestine and found that more than 20% overlap. “These data provide more compelling evidence that the brain is protected by immune cells that are educated in the gut,” explained McGavern. (NIH authors: Z. Fitzpatrick, M.L. Negro-Demontel, N. Bouladoux, D.S. Reich, Y. Belkaid, and D.B. McGavern, Nature 587:472–476, 2020; DOI:10.1038/s41586-020-2886-4)
NICHD: GENE IN MICE THAT CONTROLS FOOD CRAVINGS AND DESIRE TO EXERCISE
Scientists at NICHD have identified a gene, Prkar2a, in the mouse brain that controls the consumption of palatable, “rewarding” foods and the desire to exercise. In the absence of Prkar2a, mice reduced their intake of fatty and sugary food, even when given unlimited access to it, and were more likely to exercise. The gene’s protein product, RII-alpha, is expressed in the habenular (Hb), a tiny brain region associated with reward processing and motivation. Although the Hb’s role has been investigated in substance abuse, its role in obesity and metabolic dysregulation is unclear. RII-alpha is a component of protein kinase A (PKA), an enzyme that speeds reactions inside cells; Prkar2a deficiency decreases the PKA enzymatic activity. This effect is caused by the dysregulation of the cellular localization of the enzyme within the cell. The consequences are so pronounced that they ultimately affect the dopamine-signaling pathway. Prkar2a-deficient mice drink less of a sugar solution than wild-type mice, and the effects were strongest in males. However, Prkar2a-deficient female mice had a significant decrease in weight and metabolic efficiency and were less inclined to eat high-fat foods than males. In addition, Prkar2a-deficient mice ran 2–3 times longer on a treadmill than did wild mice. The findings from this study could provide useful information for the prevention of obesity and its accompanying risks of cardiovascular disease and diabetes. (NIH authors: E. London, J.C. Wester, M. Bloyd, S. Bettencourt, C.J. McBain, and C.A. Stratakis; JCI Insight 2020; DOI:10.1172/jci.insight.141670)
This page was last updated on Thursday, March 10, 2022
