Sandra Wolin and Ronald Germain Elected to American Academy of Arts and Sciences
IRP Scientists Recognized for RNA Revelations and Immune System Insights
Since its founding in 1780, the American Academy of Arts and Sciences has honored excellence in fields ranging from the humanities and arts to math, biology, and physics. In 2023, two of IRP’s eminent researchers joined the ranks of such luminaries as Benjamin Franklin and Nobel Prize winner Barbara McClintock as members of the Academy for their work in immunology and RNA science.
Picturing Immunity: Ronald Germain Reveals the Immune System’s Structure and Function

Dr. Ronald German’s election to the American Academy of Arts and Sciences recognized his work developing cutting-edge approaches to learning about the immune system.
IRP Distinguished Investigator Ronald N. Germain, M.D., Ph.D., realized he wanted to be an immunologist when he began working on a project for his junior high school’s science class.
“I knew where I was sitting, how old I was, and what I was looking at when I said, ‘I’m going to be an immunologist,” Dr. Germain remembers. “I didn’t want to make volcanoes or rocket ships for my science project, so I went to the library and looked through the Time Life Big Book of Science. There, I came upon a picture of a white mouse with a successfully transplanted patch of brown skin.”
Dr. Germain learned that the mouse in the book was able to tolerate the skin transplant because it was exposed to donor cells from the brown mouse in-utero, which taught the white mouse’s immune system not to attack the transplanted skin. He ended up doing a related transplantation experiment for his science fair project, setting the stage for his future career investigating the behavior of immune cells.
Today, Dr. Germain’s research laboratory at the National Institute of Allergy and Infectious Diseases (NIAID) is revealing how the spatial organization of cells influences the functioning of the immune system. By developing and using advanced imaging techniques, Dr. Germain and his research team have shed light on how immune cells move and interact within various tissues and organs in the body to cause an immune response, as well as how breakdowns in this dynamic system can lead to human disease.
To effectively marshal the body’s defenses against an infectious threat, immune cells must communicate with other types of cells, but some of those partners might be extremely hard to find amongst the trillions of cells that make up an adult human body. Dr. Germain likens it to picking up a passenger at an airport — without knowing the airline, terminal, gate, or arrival time. The likelihood of finding the passenger without any of this information is effectively zero.
“The immune system has to figure out how to solve this problem,” Dr. Germain says. “It doesn’t know what infectious agent might attack, where it will appear in the body, or when the attack will happen, but it needs to bring these immune cells together to meet the invading pathogen.”
To reveal the immune cells mustering for battle, Dr. Germain’s laboratory uses two advanced microcopy techniques to visualize cellular organization, structure, and movement. The first, called in vivo dynamic imaging, labels immune cells with fluorescent tags so his research team can watch the cells move amongst crowds of other cells and tissues in real time. Dr. Germain’s team has used this live imaging method to study animal models of human genetic diseases that affect immune cells called T cells, including a condition called X-linked lymphoproliferative disease. This serious, sometimes fatal condition, which is characterized by an extreme sensitivity to the Epstein-Barr virus, is caused by the deletion of a gene called SAP. Using its dynamic imaging tools, Dr. Germain’s lab was able to analyze T cell interactions with two other types of immune cells, respectively known as B cells and dendritic cells.
“What immediately became apparent was that the SAP-deficient T cells do not bind with B cells long enough to exchange signals and that’s why you get this defect,” Dr. Germain explains.
The techniques Dr. Germain’s lab uses can produce videos like this one, which shows T cells (red) moving amongst a structural network formed in the lymph nodes by other cells called fibroblastic reticular cells (FRC).
While this technique has long offered important insights about immune cells’ movements, it is limited to measuring only a few cell types or cellular interactions at a time. Over the last decade, however, Dr. Germain and his colleagues have developed a second, novel method to view cells in their native tissues. While the tissue and cells used in this technique aren’t alive when they are being studied, by slicing tissue into sections and labeling different molecules in the sample with different ‘antibody’ molecules, Dr. Germain’s team can build up a highly detailed and multifaceted representation of the characteristics, signaling, function, and location of many types of cells within the tissue sections. This imaging methods enables researchers to see how different types of cells interact and answer important questions like which cells inside the lymph nodes exchange the necessary information required for an immune response or how immune cells are positioned within a tumor when there is an active anti-cancer response.
Dr. Germain’s team is also using these imaging techniques to understand why the protective effects of RNA vaccines, like the ones developed for COVID-19, don’t last very long. These vaccines prompt the immune system to produce antibody molecules that mark the novel coronavirus for destruction by the immune system, but regular boosters are needed to maintain levels of those antibodies high enough to provide protection. Given that the initial doses of the vaccines produce high levels of antibodies, it’s an important mystery why antibody levels don’t remain high for years like they do after people receive vaccines that contain weakened viruses instead of just RNA, like the smallpox and yellow fever vaccines.
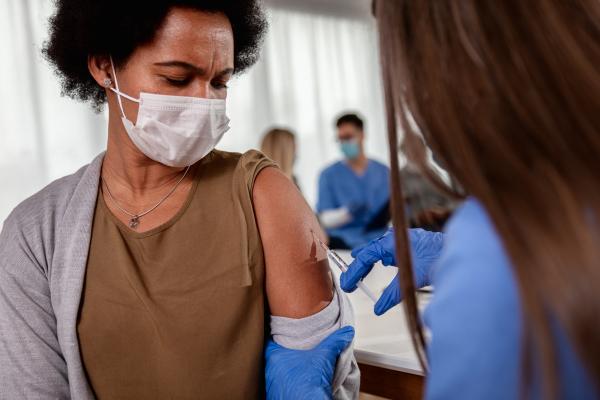
Dr. Germain’s research may one day reveal why the body requires frequent booster vaccines to maintain high levels of protection from COVID-19.
“We’re in the middle of working with companies that design vaccine adjuvants, or additional ingredients, that give a stronger, if not longer, immune response,” Dr. Germain says. “We are beginning to adapt our imaging tools to look not just in the lymphoid tissues where the immune response starts, but also in the bone marrow where it is believed that the major populations of long-lived antibody plasma cells reside. We can use our multiplex imaging tools to examine the environment that exists around these cells and our dynamic in vivo imaging to see whether the immune cells are moving in and out of these environments.”
As a hobbyist photographer, composing and analyzing images comes naturally to Dr. Germain. Fortunately for his research, the photography skills and sensibilities he has cultivated outside the lab can serve to enhance his experiments as well.
“I’m always looking for a good photograph,” he says. “That means I’m observing the environment. I’m looking for relationships and compositions. That’s what I’m doing in the lab. We try to take better and better pictures and learn more about how to analyze them to get information.”
Rounding Up Renegade RNAs: Sandra Wolin Unravels the Secrets of DNA’s Versatile Sibling

Dr. Sandra Wolin was elected to the American Academy of Arts and Sciences for her important discoveries about the ‘noncoding’ RNA molecules that carry out important tasks other than helping convert DNA’s instructions into proteins.
While scooping ice cream may sound like a scrumptious summer job to some, IRP senior investigator Sandra Wolin, M.D., Ph.D., was ready for a change when she began her freshman year at Princeton University. After visiting department after department looking for a position, she found a home tending fruit flies for one of the university’s biology labs.
“They started giving me little projects and experiments to do, and I got hooked,” Dr. Wolin says.
Soon the ‘Freshman Fruit Fly Flipper,’ as her colleagues nicknamed her, became a biochemistry major, setting the stage for a distinguished career in genetics and RNA biology. Her research at both her previous laboratory at Yale University and in the IRP, which she joined in 2017, has focused on making sense of the structure, function, and biological roles of RNA and finding ways to apply this knowledge to new diagnostics and therapies for human diseases.
RNA — short for ribonucleic acid — is a vital molecule made of a single strand of nucleic acids, the same building blocks that make up double-stranded DNA. RNA’s best-known function is delivering instructions from the DNA in a cell’s nucleus to the protein manufacturing machinery elsewhere in the cell. However, most RNA is actually so-called ‘noncoding RNA,’ which carries out a multitude of other necessary cellular tasks, including repairing the ends of chromosomes, synthesizing proteins, and helping to bring newly made proteins to their proper destinations in the cell. These RNAs often must join with proteins to form ‘RNA-protein complexes’ in order to perform these functions.
Dr. Wolin’s laboratory studies noncoding RNAs and their protein partners to understand their breadth of functions within the cell. In addition, she examines how cells recognize and remove damaged or defective strands of noncoding RNA, as well as the consequences when damaged RNAs stick around.
“This is important to human health because some of these RNA-protein complexes are targets for autoantibodies in diseases like systemic lupus erythematosus and other autoimmune conditions,” Dr. Wolin says.

While DNA is confined to a cell’s nucleus and mitochondria, RNA is free to roam around the cell, allowing to contribute to a wide array of cellular functions.
Using genetic and biochemical analyses of cells from bacteria, mice, and humans, Dr. Wolin’s research team is determining the roles of various RNA-protein complexes that have been associated with autoimmune reactions. By identifying the same molecules across different organisms, Dr. Wolin’s team can transfer insights from one type of organism to another to build a bigger picture of what is happening and hone in on the mechanisms causing the problem.
One of Dr. Wolin’s proudest achievements was the identification and characterization of a new way that cells dispose of unwanted RNA. Her laboratory showed that when a ring-shaped protein called Ro60 binds to the stem of a lollipop-shaped noncoding ‘Y RNA,’ the resulting Ro60 ribonucleoprotein (RNP) can then bind to another enzyme to form a cellular machine that breaks down unneeded RNA molecules for disposal. What’s more, Dr. Wolin’s lab discovered how this RNA shredder works: the unwanted RNAs thread through the ring-shaped Ro60 molecule, which helps unfold the RNAs and facilitates their degradation by the enzyme.
“This new RNA degradation machine that we found represents a new role for noncoding RNAs,” Dr. Wolin says.
On the other hand, RNA–protein complexes sometimes become targets for people’s immune systems, leading to ‘autoimmune’ diseases in which the immune system attacks the body’s own tissues. For example, long before symptoms emerge in patients with the autoimmune condition systemic lupus erythematosus (SLE), doctors can detect ‘autoantibody ‘molecules that cause the immune system to attack R060 RNPs. Intriguingly, Dr. Wolin’s laboratory has also discovered that some bacteria that normally inhabit our bodies have their own Ro60 RNPs. This has led to the theory that, in some SLE patients, the autoantibodies might start out as antibodies that target the bacterial Ro60 RNPs before expanding their list of targets to include the Ro60 RNPs in patients’ own cells.
There is also growing evidence that the presence of excess noncoding RNAs, particularly mutated or otherwise defective RNAs, may predispose people to autoimmune diseases by triggering excess production of the immune chemical interferon. Dr. Wolin hypothesizes that this may contribute to the formation of some autoantibodies in patients.
“Understanding how cells recognize problematic RNAs and dispose of them may eventually help us prevent synthesis of interferon and treat autoimmune disease,” Dr. Wolin explains.

Dr. Wolin celebrates her election to the American Academy of Arts and Sciences with members of her lab.
With the abundant resources of NIH at her disposal, Dr. Wolin is optimistic about her chances of achieving those kinds of advances.
“What I love about being here is that I get to do discovery science,” Dr. Wolin says. “In the past, I had to write a grant and tell people what I was going to find before I could get funding to do the experiments. Now I can just follow my nose to see where the data takes us, and that’s what I’m going to keep doing.”
Subscribe to our weekly newsletter to stay up-to-date on the latest breakthroughs in the NIH Intramural Research Program.
Related Blog Posts
This page was last updated on Wednesday, September 6, 2023
