Reducing Stress Boosts Efficiency of Bacterial Factories
Unconventional Genetic Strategy Could Enhance Production of Medical Treatments
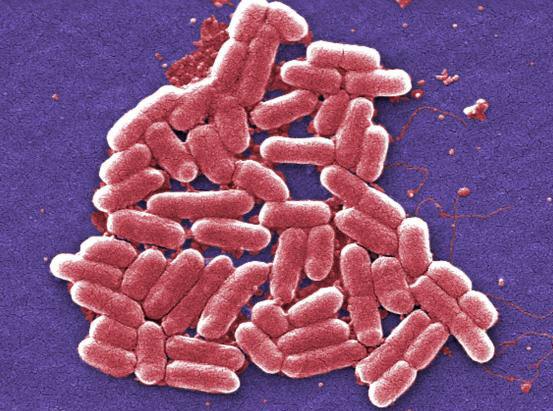
Image credit: Janice Haney Carr, U.S. Centers for Disease Control and Prevention
Recent IRP research details a new strategy that could make genetically modified E. coli bacteria, and possibly other microbes, better at producing medicines and other important substances.
We all have bad days on the job — your colleague keeps bugging you, your boss yelled at you for an innocent mistake, and you skipped lunch because you have five different deadlines coming up. Understandably, many people find it much harder to get their work done under such stressful circumstances. Microbes that produce chemicals for medicine and scientific research experience similar struggles, but a recent IRP study has found that short-circuiting their stress response makes them far more efficient at that task.1
Single-celled organisms like fungi and bacteria produce many substances used for medical treatment or as key ingredients in vaccines, as well as molecules that scientists need for their research. To get microbes to manufacture these products, known as biologics, scientists insert genes into them that cause them to make the chemicals. Over time, researchers have found ways to get each microbial cell to produce more of these molecules, whether by optimizing the conditions in which the cells are grown or through genetic manipulations that rev up the cellular systems involved in protein synthesis.
Perhaps unsurprisingly, microbes don’t particularly like it when we hijack their protein production systems. In fact, forcing microorganisms to make a substance they would not normally produce triggers a stress response that impairs both their ability to grow and their capacity to carry on their molecular manufacturing. This occurs because stress switches on certain genes that suppress those processes, which means that the conventional approach of making specific genes more active to boost yields would not solve the problem. Instead, researchers led by IRP senior investigator Joseph Shiloach, Ph.D., aimed to short-circuit the stress response entirely by suppressing a few key genes that become more active when microbial cells experience stress.
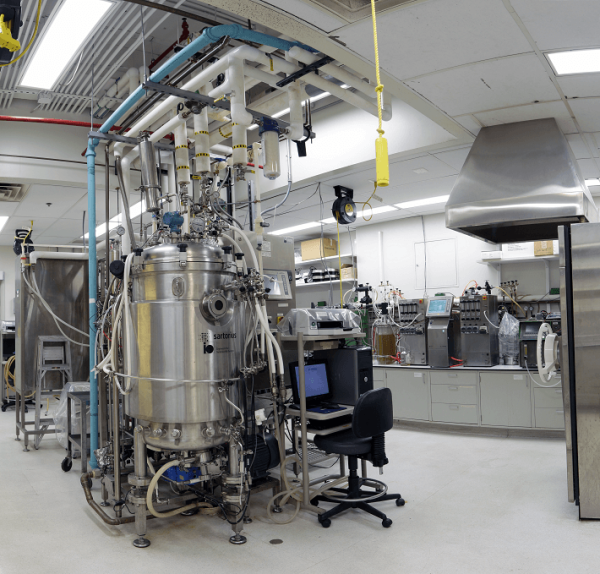

Dr. Shiloach’s team grows a variety of microbial and other types of cells in fermentation tanks like this one in order to figure out ways to make them better at synthesizing useful molecules.
“Traditionally, scientists have only looked at genes that become less active when growth slows down, and there are many genes like that, but it’s hard to improve the activity of all of those genes because there are so many,” says Ashish Sharma, Ph.D., a postdoctoral research fellow in Dr. Shiloach’s lab and the new study’s first author. “With our systems-level approach, we do just one or two genetic modifications and we get much better production.”
The IRP scientists tested this unconventional strategy in E. coli bacteria. Decades of research have produced detailed maps of how all the different genes in E. coli influence one another, as well as which genes become more or less active when E. coli are forced to produce substances they don’t naturally make. This knowledge allowed the IRP team to focus only on genes that become more active in that situation and that do not regulate other genes, which makes the consequences of manipulating them easier to track. The researchers examined the effects of deleting each one of these genes individually in E. coli that had been genetically engineered to produce a glowing green protein called green fluorescent protein (GFP), and they discovered that eliminating many of these genes dramatically boosted the amount of GFP each bacterial cell produced, sometimes by more than 250 percent.
More From the IRP
Blog
Mouth Microbes Turn Treasonous in Gum Disease
In a second set of experiments, the IRP team took a list of the ten single-gene deletions that boosted yields the most and created additional strains of E. coli that each lacked a different pair of those genes, for a total of 45 different combinations. They repeated that experiment twice: once with E. coli designed to make GFP and once with E. coli designed to make a more complex molecule called L-asparaginase that is used to treat acute lymphoblastic leukemia, a type of blood and bone marrow cancer. Some of the combinations of gene deletions increased yields more than any of the single-gene deletions, although others actually hindered production. Seeing higher yields of L-asparaginase was particularly good news because, unlike GFP, L-asparaginase is secreted from cells after it is made rather than building up inside them.

Dr. Sharma (left) and Dr. Shiloach in their lab.
“The cell has limited space inside it, so if something just accumulates inside the cell, eventually there’s no space left,” explains Dr. Sharma, “but if the modified bacteria over-produce proteins that are being exported out of the cell, then that shows the real ability of the cell to produce them.”
By combining this new strategy with existing practices for boosting microbial protein production, researchers could noticeably enhance the ability of E. coli to manufacture a wide array of medicines and chemicals for scientific and industrial use, thereby speeding up the process and lowering costs. Moreover, if scientists can identify the genes that trigger the stress response in mammalian cells, they could also apply the IRP researchers’ approach to boost production of the particularly complex molecules those cells are used to synthesize.
“Eventually, if you can come up with a cell that is a better producer, you can use it every time you need to produce a protein to see if you can get higher production,” says Dr. Shiloach. “You have a better producer in general.”
Subscribe to our weekly newsletter to stay up-to-date on the latest breakthroughs in the NIH Intramural Research Program.
References:
[1] A novel knock out strategy to enhance recombinant protein expression in Escherichia coli. Sharma AK, Shukla E, Janoti DS, Mukherjee KJ, Shiloach J. Microb Cell Fact. 2020 Jul 23;19(1):148. doi: 10.1186/s12934-020-01407-z.
Related Blog Posts
This page was last updated on Tuesday, January 30, 2024
