Research Briefs
Read about NIH scientific advances and discoveries by intramural scientists:
- NEI: PROTEIN FRAGMENTS PROTECT AND STIMULATE RETINAL NEURONS
- NCI: INTERNATIONAL STUDY FINDS GENETIC RISK FACTORS FOR RARE CHILDHOOD CANCER
- NHGRI, NIEHS: MANY PATIENTS CHANGE THEIR MIND ABOUT RECEIVING SECONDARY GENOMIC FINDINGS
- NIAID, NCI: ANCIENT VIRUSES, BACTERIA, AND HOST IMMUNE SYSTEM PARTICIPATE IN “MULTIKINGDOM DIALGOUE”
- NINDS, NIMH: TAKING SHORT BREAKS MAY HELP BRAIN LEARN NEW SKILLS
- NIAID: ANTIBODY EFFECTIVE AGAINST MALARIA INFECTION
NEI: PROTEIN FRAGMENTS PROTECT AND STIMULATE RETINAL NEURONS
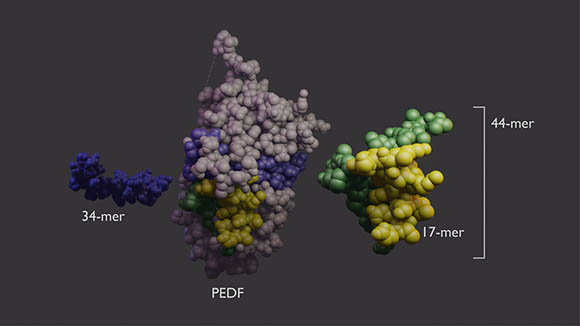
CREDIT: LESLEY EARL, NEI
PEDF protein (center) has two domains with different functions. The 34-mer (blue, left) has anti-angiogenic properties. The 44-mer (green and yellow, right) protects and stimulates neurons. The 17-mer (yellow) is a smaller region of the 44-mer with the same function.
In an NEI-led study, researchers found that small protein fragments known as peptides serve distinct functions in protecting neurons found in the light-sensing layer of the retina. These peptides are components of pigment epithelium-derived factor (PEDF), which prevents cell death as well as the invasion of blood vessels and inflammation, hallmarks of eye diseases such as age-related macular degeneration, retinitis pigmentosa, and diabetic retinopathy.
In this study, investigators cultured immature rat retinal cells using an in vitro model. These cells were isolated in a nutrient-deprived environment in which researchers tested the protective function of each PEDF fragment. They discovered that two closely associated peptides (44-mer and 17-mer) prevented premature cell death and promoted the growth of new connections between neurons. One other peptide known as 34-mer lacked such retinal protective property, but it is known for stopping abnormal blood vessel growth in the eye, as shown in a previous study.
The findings provide more detail of the mechanisms behind PEDF and could lead to new treatments for degenerative eye diseases. “We’re hoping that we can harness some of these protective effects in a peptide-based therapeutic approach in the near future,” said lead author Patricia Becerra. (NIH authors: G. Michelis, R. Villasmil, and S.P. Becerra, J Neurochem 2021; DOI:10.1111/jnc.15454)
NCI: INTERNATIONAL STUDY FINDS GENETIC RISK FACTORS FOR RARE CHILDHOOD CANCER
An international consortium of scientists, led by the NCI, has discovered new genetic risk factors for rhabdomyosarcoma (RMS), a rare cancer that forms in soft tissues and most often affects children. RMS treatment has historically been guided by clinical factors such as tumor size, location in the body, and the presence of PAX-FOX01, a fusion gene associated with poorer survival. These new findings emerge from the largest genomic profiling effort of RMS tumors to date and establish links between genetic risk and clinical outcomes so that treatment can be better tailored to the individual.
The researchers used next-generation sequencing to analyze 641 tumor samples from children with RMS. The results revealed that patients with mutations in the genes TP53, MYOD1, and CDKN2A had worse outcomes than children without these mutations. Moreover, the investigators identified a median of one mutation per tumor and found that patients with two or more mutations per tumor might also have poorer survival outcomes.
“These discoveries change what we do with these patients and trigger a lot of really important research into developing new therapies that target these mutations,” said NCI’s Javed Khan, who led the study. (NIH authors: J.F. Shern, R. Patidar, H-C. Chou, Y.K. Song, M.E. Yohe, S. Sindiri, J. Wei, X. Wen, K. Jones, B. Hicks, and J. Khan, J Clin Oncol 2021; DOI:10.1200/JCO.20.03060)
NHGRI, NIEHS: MANY PATIENTS CHANGE THEIR MIND ABOUT RECEIVING SECONDARY GENOMIC FINDINGS
A small minority of patients undergoing genome sequencing elect not to know whether their results uncover any secondary genetic findings (SFs). SFs are known, actionable DNA variants associated with serious diseases that are unrelated to the primary reason for an individual’s medical care or participation in a study. “Because these genomic findings can have life-saving implications, we wanted to ask the question: Are people really understanding what they are saying ‘no’ to?” said lead author Benjamin Berkman. “If they get more context, or a second opportunity to decide, do they change their mind?”
The investigators recruited 231 participants from a large NIEHS study who initially accepted or refused to be notified of any SFs that came out of their genome sequencing. Each respondent completed an online survey that asked questions about their decision-making process and included an intervention–more detailed information about SFs and the option to change their original decision: Nearly half of the initial refusers changed their mind. Surprisingly, 45.8% of refusers thought they had initially agreed to receiving SFs (compared with 6.8% of acceptors who thought they had refused). Among the persistent refusers, the most common reason for refusing was concern about becoming worried or sad. Based on these results, the researchers suggest considering a transparent default of returning secondary findings without soliciting participant preferences. Alternatively, subjects could be provided with detailed information about SFs and allowed multiple opportunities to revise their decision. (NIH authors: W. Schupmann, S.A. Miner, J.E. Hall, S.H. Schurman, and B.E. Berkman, Genet Med 2021; DOI:10.1038/s41436-021-01271-1)
NIAID, NCI: ANCIENT VIRUSES, BACTERIA, AND HOST IMMUNE SYSTEM PARTICIPATE IN “MULTIKINGDOM DIALGOUE”
CREDIT: NHGRI
The microbiome is comprised of microorganisms that live in and on us and contribute to human health and disease.
Collectively known as microbiota, billions of microbes inhabit the human body and play an important role in immunity, digestion, and other aspects of host physiology. This symbiotic relationship extends not just to microbes such as bacteria and fungi, but to endogenous retroviruses (ERVs)—remnants of viruses that infected our ancestors and became integrated into the human genome. In a NIH-led study, scientists found that some of these ERVs mediate an alliance between the microbiota and the host immune system.
In mouse and in vitro experiments, the researchers discovered that the common skin bacterium Staphylococcus epidermidis promoted expression of several ERVs in skin cells known as keratinocytes. This response was associated with maintaining protective tissue immunity and promoting tissue healing. These beneficial processes were impaired when the scientists inhibited expression of ERVs, further demonstrating a retrovirus-facilitated communication between the microbiota and skin immune system.
Conversely, a high-fat diet in the mouse model triggered an inflammatory response to the microbiota. This pro-inflammatory reaction was controlled with an antiretroviral treatment, suggesting that certain nutritional conditions can change how ERVs communicate with the microbiota. The findings shed new light on the mechanisms behind diseases such as obesity and inflammatory skin disorders. “Together,” the authors wrote, “our results support the idea that the host may have co-opted its endogenous virome as a means to communicate with the exogenous microbiota, resulting in a multi-kingdom dialog that controls both tissue homeostasis and inflammation.” (NIH authors: D.S. Lima-Junior, S.R. Krishnamurthy, N. Bouladoux, N. Collins, S. Han, M.G. Constantinides, V.M. Link, A.I. Lim, M. Enamorado, C. Cataisson, L. Gil, I. Rao, T.K. Farley, G. Koroleva, S.H. Yuspa, and Y. Belkaid, Cell 184:3794–3811.e19, 2021; DOI:10.1016/j.cell.2021.05.020)
NINDS, NIMH: TAKING SHORT BREAKS MAY HELP BRAIN LEARN NEW SKILLS
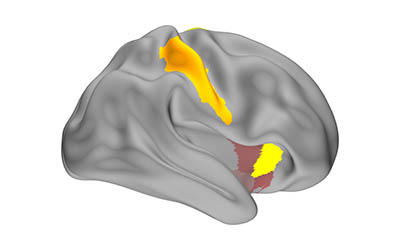
CREDIT: COHEN LAB, NINDS
In a study of healthy volunteers, NIH researchers discovered that our brains may replay compressed memories of learning new skills when we rest. Above is a map of the memory replay activity observed in the study.
In a recent study, NIH scientists explored how wakeful conversion of learned experiences to long-term memory was reinforced by neural replay, a memory-consolidating process in which the brain automatically replays a skill in a compressed time frame. The NINDS-led team found that skill improvements were largely explained by substantial jumps in performance after a brief period of rest between practice sessions, as opposed to improvements during actual practice.
Investigators asked subjects to type a five-digit numeric sequence on a response pad as many times as they could with their nondominant hand. A 10-second practice session was followed by a 10-second rest period and repeated 36 times. To analyze how each subject processed the new skill, researchers measured brain electrical activity using magnetoencephalography.
They found that neural replay of the skill during the rest interval was temporally compressed by approximately 20-fold relative to the practiced behavior. On average, subjects replayed the sequence about six times more than they practiced it. Furthermore, subjects who replayed the sequence more often during rest periods performed the sequence faster in the following practice session compared with those who replayed it less frequently. The findings suggest that taking structured breaks may aid in learning new skills. “Overall,” said senior author Leonardo Cohen, “our results support the idea that manipulating replay activity during waking rest may be a powerful tool that researchers can use to help individuals learn new skills faster and possibly facilitate rehabilitation from stroke.” (NIH authors: E.R. Buch, L. Claudino, R. Quentin, M. Bönstrup, and L.G. Cohen, Cell Reports 10:109193, 2021; DOI:10.1016/j.celrep.2021.109193)
NIAID: ANTIBODY EFFECTIVE AGAINST MALARIA INFECTION
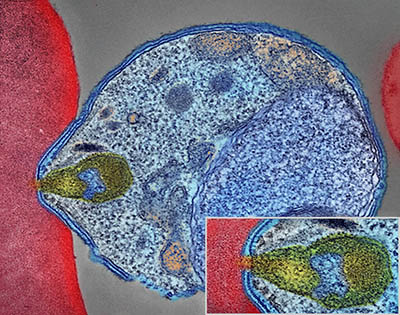
CREDIT: NIAID
Colorized electron micrograph showing malaria parasite (right, blue) attaching to a human red blood cell. The inset shows a detail of the attachment point at higher magnification.
A recent small clinical trial led by NIAID scientists found that a single dose of a monoclonal antibody safely prevents malaria infection for up to nine months. Malaria infection occurs when mosquitos infected with the plasmodium parasite inject an immature form of the parasite known as sporozoites into the bloodstream. These sporozoites travel to the liver to mature and multiply, ultimately resulting in severe illness or death. The new laboratory-produced antibody, called CIS43LS, was developed to remain present within the bloodstream over time and neutralize sporozoites before they invade the liver.
Investigators first administered one dose of CIS43LS to 21 healthy subjects by either intravenous infusion or subcutaneous injection, with each dose varying between 5 and 40 milligrams per kilogram of body weight. Over the next six months participants were followed to ensure that treatment was well tolerated and that plasma antibody concentrations remained durable.
In the second part of this study, 15 participants (nine who had received CIS43LS and six control participants) were subjected to a carefully monitored exposure to malaria from infected mosquitoes. None of the volunteers receiving the antibody developed malaria, whereas five of the six control participants did (and were promptly treated to eliminate the infection). Among those who received an infusion of CIS43LS did so at time points ranging from 1 to 9 months before being exposed to malaria, demonstrating how antibody treatment could be a safe and effective way to prevent the disease. A larger phase 2 clinical trial is currently underway in Mali to evaluate CIS43LS in adults living under malaria endemic conditions. (NIH authors: M.R. Gaudinski, N.M. Berkowitz, A.H. Idris, E.E. Coates, L.S.A. Holman, F. Mendoza, I.J. Gordon, S.H. Plummer, O. Trofymenko, Z. Hu, S. O’Connell, M. Basappa, N. Douek, S.R. Narpala, C.R. Barry, A.T. Widge, R. Hicks, S.F. Awan, R.L. Wu, S. Hickman, D. Wycuff, J.A. Stein, C. Case, K. Carlton, J.G. Gall, S. Vazquez, B. Flach, G.L. Chen, J.R. Francica, B.J. Flynn, N.K. Kisalu, A. McDermott, J.R. Mascola, J.E. Ledgerwood, and R.A. Seder for the VRC 612 Study Team, N Engl J Med 2021; DOI:10.1056/NEJMoa2034031)
This page was last updated on Tuesday, February 1, 2022
