New Methods
ScPopCorn Gets to the Kernel of Single-Cell Experiments
NLM–Made Algorithm Helps NIH Researchers
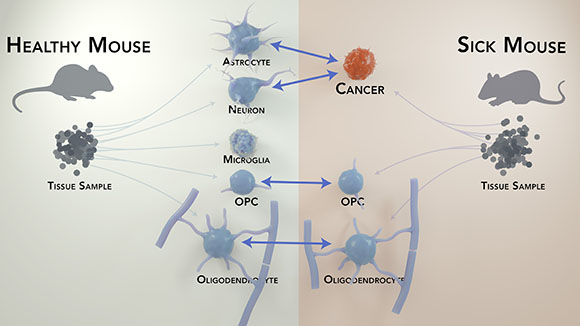
CREDIT: DONALD BLISS, NLM
A new computational method, scPopcorn, allows identification of individual cells by determining which genes are being expressed and can compare them with the same cells types in other animals.
Researchers at the National Library of Medicine’s National Center for Biotechnology Information have created a new algorithm called scPopCorn (single-cell subpopulations comparison) to capture the differences among populations of cells from single-cell experiments. The algorithm, developed by my team is available at GitHub and is described in an article in Cell Systems (Y. Wang, J. Honka, and T.M. Przytycka, Cell Syst 8:506–513, 2019).
The most frequently performed analyses of single-cell RNA sequencing (scRNA-seq) datasets include the identification of subpopulations of cells in scRNA-seq experiments and the comparison of such subpopulations across experiments. In multicellular organisms, different cell types execute different transcriptional programs expressing different sets of genes. Current experimental techniques can measure gene expression at single-cell resolution, making it possible to address questions that could not be answered with standard bulk experiments in which the total gene expression from a heterogeneous cell population was measured.
Single-cell transcriptomics opens a window to a better understanding of changes in the functioning of cell populations across different states and conditions including diseases. However, new computational methods are required to effectively gain important insights from these, unfortunately still quite noisy, measurements.
To address this need, my team leveraged several new algorithmic ideas and introduced the computational method scPopCorn. Unlike previous methods that treated the identification of cell types and their comparison across experiments as two separate tasks, scPopCorn identifies subpopulations of cells in individual experiments simultaneously by incorporating these two tasks into one complex optimization problem.
The optimization involves a measure of the homogeneity of a cell population (population consistency), which when combined with a technique much like Google’s personalized PageRank approach, guides subpopulation detection. (PageRank is Google’s algorithm that ranks web pages in search engine results.)
In addition, a cell-to-cell similarity measure is used to guide the mapping. In the scPopCorn method, the researchers substituted a cell-to-cell expression similarity graph for the network of webpages, and for each cell, estimated its preference (a “vote”) for which other cells should be included in the same subpopulation.
This integrative approach helps researchers confidently define both the common and unique cell types across many experiments. Scientists can use this method to understand and map the differences among populations of cells with different disease status and developmental stages and of different sexes and species. In particular, scientists can use the algorithm to identify similar and distinct cell types present in such single-cell experiments.
This new computational method, scPopCorn, not only enabled the design of a highly accurate identification of subpopulations and a mapping approach, but also introduced mathematical concepts that can serve as stepping stones for other tools to interrogate the relationships among single cells.
(NIH authors: Y. Wang, J. Hoinka, and T.M. Przytycka, Cell Syst 8:506–513, 2019; DOI:10.1016/j.cels.2019.05.007)
This page was last updated on Wednesday, March 30, 2022
