Research Briefs
NEI: STEM-CELL THERAPY PREVENTS BLINDNESS IN ANIMAL MODELS OF RETINAL DEGENERATION

CREDIT: NEI
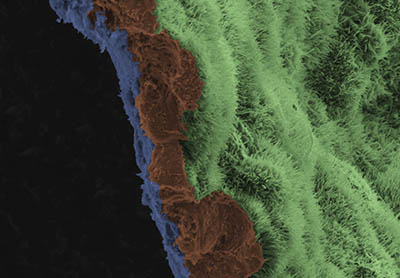
A scanning-electron-microscope image shows a polarized retinal pigment epithelial (RPE) cell monolayer on a biodegradable scaffold. The image is colored to highlight the scaffold in blue, and three RPE cells in brown. The apical process of cells in RPE monolayer are light green.
Researchers at NEI used a novel stem-cell-based therapy to prevent blindness in rat and pig models of geographic atrophy, the advanced “dry” form of age-related macular degeneration (AMD). Specifically, induced pluripotent stem cells (iPSCs), which can become any cell type in the body, were derived from the blood cells of three human AMD patients. The iPSCs were programmed to become retinal pigment epithelial (RPE) cells, which nurture light-sensing cells, called photoreceptors, in the retina. During the progression of AMD, RPE cells often die, which leads to the death of the photoreceptors and thus to blindness. In the study, the iPSC-derived RPE cells were grown as patches one-cell thick—on a biodegradable scaffold—to mimic their natural structure within the eye. After the RPE patches were transplanted into normal and laser-induced degenerating retinas of animals, imaging studies showed that the cells integrated into the retina. Additionally, immunostaining for RPE65, a gene expressed by mature RPE cells that is essential for vision, confirmed that the transplanted iPSC-derived RPEs were able to support and safeguard photoreceptors. Although a concern with stem-cell therapies is the ability of transplanted stem cells to acquire oncogenic mutations that lead to uncontrolled growth, the researchers found that iPSC-derived RPE cells had no mutations linked to cancer in an oncogene analysis. This study provides preclinical evidence of the safety and efficacy of iPSC-derived RPE cells as a treatment for AMD and a framework for the translating iPSC therapies into clinical trials. (NIH authors: R. Sharma, V. Khristov, A. Rising, B.S. Jha, R. Dejene, N. Hotaling, Y. Li, Q. Wan, C. Zhang, M.M. Campos, K.J. Miyagishima, D. McGaughey, R. Villasmil, M. Mattapallil, H. Qian, W. Wong, S. Miller, A. Marminishkis, J. Amaral, and K. Bharti, Sci Transl Med 11:eaat5580, 2019; DOI:10.1126/scitranslmed.aat5580)
NCI, NLM: ARTIFICIAL INTELLIGENCE OUTPERFORMED HUMANS IN IDENTIFYING CERVICAL PRECANCER
Investigators at NCI, NLM, and Global Good (Bellevue, Washington) developed an artificial-intelligence approach, called automated visual evaluation, that can analyze digital images of a woman’s cervix and accurately identify cervical precancers. More than 60,000 digitized cervical images from a cervical-cancer screening study were used to train a deep-learning algorithm to recognize patterns of precancerous changes. When this computational precancer detection approach was compared with human experts in the evaluation of Pap tests, a widely used cervical-cancer screening approach, the algorithm was better at identifying precancer. Moreover, this new method could potentially improve cervical-cancer screening in low-resource settings where cervical-cancer is a leading cause of death. Health workers in these types of settings need only minimal training to use a cell phone or camera for cervical screening and treatment. The investigators are planning to use a wide variety of cameras to further train the algorithm on images of normal and precancerous cervical tissue that are collected from women around the world. These studies are important to identify subtle differences in the appearance of the cervix and precancer among women in different geographical areas. The goal of this research is to create an optimal artificial-intelligence-based screening method for cervical precancer that is publicly available for use and can be used to improve the control of cervical cancer worldwide. (NIH authors: S. Antani, Z. Xue, K. Yu, L.R. Long, M. Demarco, J.C. Gage, A.C. Rodriguez [consultant to NCI], N. Wentzensen, and M. Schiffman, J Natl Cancer Inst 2019; DOI:10.1093/jnci/djy225)
NIAID (VRC): INVESTIGATIONAL MONOCLONAL ANTIBODY TO TREAT EBOLA IS SAFE
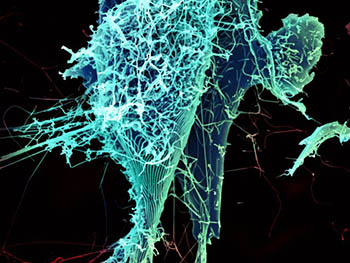
CREDIT: H. FELDMANN, P. JAHRLING, E. FISCHER, AND A. MORA, NIAID
After multiplying inside a host cell, the stringlike Ebola virus is emerging to infect more cells. Ebola is a rare, often fatal disease that occurs primarily in tropical regions of sub-Saharan Africa.
In May 2018, NIAID investigators at the institute’s Vaccine Research Center (VRC) and their collaborators began a phase 1 clinical trial examining the safety and efficacy of an investigational Ebola treatment with the monoclonal antibody mAb114. The trial revealed that intravenous mAb114 is safe, well-tolerated, and easy to administer. The participants in the trial, which was conducted at the NIH Clinical Center, were 18 healthy human volunteers. The treatment is also being offered to Ebola patients in the Democratic Republic of Congo (DRC) under compassionate use and as a phase 2/3 clinical trial of multiple investigational treatments.
The mAb114 binds to the core-receptor-binding domain of the Ebola virus glycoprotein, preventing the virus from infecting human cells. Until the discovery of mAb114—which scientists derived from antibodies isolated from the blood of a survivor from the 1995 Ebola outbreak in DRC—there had been no molecules reported to target this well-hidden core. Studies showed that mAb114 was able to protect nonhuman primates from lethal Ebola virus when given as late as five days after infection. The VRC developed mAb114 in collaboration with scientists in the DRC, Switzerland, and the U.S. Army Medical Research Institute of Infectious Diseases at Fort Detrick, Maryland. (NIH authors: M.R. Gaudinski, E.E. Coates, L. Novik, A. Widge, K.V. Houser, E. Burch, L.A. Holman, I.J. Gordon, G.L. Chen, C. Carter, M. Nason, S. Sitar, G. Yamshchikov, N. Berkowitz, C. Andrews, S. Vazquez, C. Laurencot, J. Misasi, F. Arnold, K. Carlton, H. Lawlor, J. Gall, R.T. Bailer, A. McDermott, R.A. Koup, J.R. Mascola, B.S. Graham, N.J. Sullivan, and J.E. Ledgerwood, Lancet 2019; DOI:10.1016/S0140-6736(19)30036-4)
NIAID (ROCKY MOUNTAIN LABS): TICK SALIVARY GLANDS MAY HOLD SECRET FOR HOW VIRUSES ARE TRANSMITTED
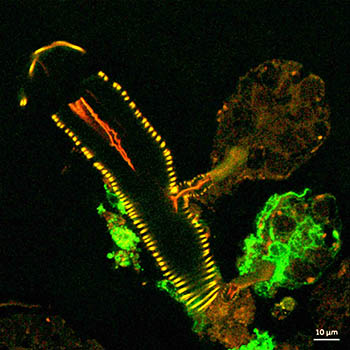
CREDIT: NIAID
This confocal microscope image shows a cross-section of a tick salivary gland infected with Langat virus (green). Two rounded structures on the right, called acini, are shown attached to a duct (yellow). The lower acinus is infected, as denoted by the green fluorescent signal.
The salivary glands of Ixodes scapularis (more commonly known as black-legged ticks or deer ticks) may hold the secret to preventing tick-borne infections. Tick-borne flaviviruses (TBFVs) have been on the rise in North America for the last two decades and cause roughly 15,000 infections each year. Flaviviruses include dengue virus, Zika virus, West Nile virus, yellow fever virus, and, in North America, the Powassan virus (POWV), which occurs primarily in northeastern states and the Great Lakes region.
Tick bites are especially dangerous because the TBFVs are so rapidly transmitted to the host. In just 15 minutes, an infected tick can transmit POWV to a human host or other mammal, causing long-term neurological disease and even death in 10 percent of those infected. TBFVs are found in the salivary glands (SGs) of ticks and are believed to be transmitted in the early stages of feeding.
Researchers at NIAID’s Rocky Mountain Labs (Hamilton, Montana) found, in a 2017 study, that tick SGs could be cultured and studied ex vivo to determine how TBFVs multiplied. The new study builds on those findings to compare the SGs of fed (fully engorged) ticks with those of unfed ticks to evaluate SG cultures and virus-transmission pathways. The research showed that the concentrations of TBFVs were higher in the SGs of fed ticks and that only certain kinds of SGs are infected. Additionally, the scientists identified a specific tick gene that is involved in infection. These findings might point to transmission pathways that could potentially be blocked and open doors to new methods of preventing infections. (NIH authors: J.M. Grabowski, O.R. Nilsson, E.R. Fischer, D. Long, D.K. Offerdahl, D.P. Scott, and M.E. Bloom, mBio 2019; DOI:10.1128/mBio.02628-18)
NINDS: NEURODEGENERATIVE DISORDERS MAY BE TRIGGERED BY HYPERACTIVE IMMUNE RESPONSE
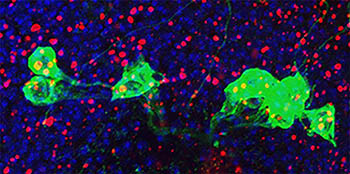
CREDIT: E. GINIGER LAB, NINDS
NINDS scientists found that, in fruit flies, the overexpression of the Cdk5-alpha gene caused the cell’s waste system (red) to back up and trigger the immune system to attack dopamine neurons (green and blue).
The human body is a complex system of checks and balances, but exactly how systems affect one another isn’t always clear. Researchers at NINDS found, in experiments using fruit flies (Drosophila melanogaster), that the body’s immune system may play a critical role in the damage caused by aging brain disorders. The study involved altering the activity of the Cdk5 protein kinase, a group of proteins that are important in early brain development and may be involved in neurodegenerative diseases (such as Parkinson and Alzheimer diseases), degeneration of neurons, impaired autophagy, and accelerated aging. The researchers showed that increased activity of Cdk5 slowed autophagy (a process that rids the body of damaged cells) and starts a chain reaction that triggers an increase in the innate immune response, which in turn causes the immune system to attack and kill the body’s own neurons. The researchers suspect that this destructive chain reaction may occur during several neurodegenerative disorders. These conclusions provide a new way of looking at the progression of neurodegenerative diseases and could offer alternative solutions for stopping or slowing them. (NIH authors: A.K. Shukla, J. Spurrier, I. Kuzina, and E. Giniger, Cell Rep 26:131–144.e4, 2019; DOI:10.1016/j.celrep.2018.12.025)
NIEHS, NHGRI, NTP: LIFE EXPERIENCES MAY ALTER DNA METHYLATION
According to scientists at NIEHS, NHGRI, and the interagency National Toxicology Program (NTP), DNA sequence may determine where methylation events occur, but life events, such as pregnancy, could alter these DNA methylation patterns. The researchers used inbred mice to examine methylation, the addition of methyl groups to DNA, in regions of DNA where the nucleotide cytosine is followed by a phosphate and the nucleotide guanine. Known as CpG sites, these areas of methylation are the most common form of DNA modification in mammals and, based on this research, may serve as an epigenetic record of life events in body cells.
Researchers intercrossed two inbred mouse strains (C57BL/6N and C3H/HeN) and measured DNA-methylation patterns in the livers of parents and progeny using whole-genome bisulfite sequencing. They found that DNA methylation patterns were closely linked to the genotype of an animal and were passed to male and female offspring unchanged. However, female animals that had experienced pregnancy—compared with virgin females—had hundreds of sites in their genome where methylation was decreased. These findings suggest that the quality of binding between some transcription factors and their DNA targets is influenced by genetics, resulting in different DNA methylation patterns. Furthermore, methylation pattern changes observed in females that had given birth suggest that major life events may leave a distinct methylation pattern signature in the genome.
While the team has not determined exactly how these female mice incurred a loss of DNA methylation, the work bolsters the importance of epigenetics in influencing gene regulation. (NIH authors: S.A. Grimm, T. Shimbo, M. Takaku, J.W. Thomas, S Auerbach, B.D. Bennett, J.R. Bucher, A.B. Burkholder, F. Day, Y. Du, C.G. Duncan, J.E. French, J.F. Foley, J. Li, B.A. Merrick, R.R. Tice, T. Wang, X. Xu, NISC, NHGRI Comparative Sequencing Program, P.R. Bushel, D.C. Fargo, J.C. Mullikin, and P.A. Wade, Nat Commun 10:305, 2019; DOI:10.1038/s41467-018-08067-z)
NIAID: WITH HIV, UNDETECTABLE EQUALS UNTRANSMITTABLE

CREDIT: NIAID
Antiretroviral drugs to treat HIV infection spill out of a pill bottle.
The HIV undetectable = untransmittable (U=U) concept has been validated by NIAID scientists who reviewed and summarized results from large clinical trials and cohort studies. The U=U concept suggests that people infected with HIV who achieve and maintain an undetectable viral load by taking antiretroviral therapy (ART) cannot sexually transmit the virus to others. The studies showed that no HIV transmission occurred in heterosexual and male-male couples, in which one partner is uninfected and the other has HIV and a durably suppressed viral load. For U=U to work as an HIV-prevention strategy, the undetectable viral load must be sustained by taking ART daily as prescribed. The validation of this concept has many behavioral, social, and legal implications and can help change the stigma surrounding the disease. The authors point out, however, that ART adherence may be difficult when those infected with HIV don’t have access to quality health care and that it’s important to implement programs that help patients remain in care and to address the challenges in their lives that result in their stopping therapy. (NIH authors: R.W. Eisinger, C.W. Dieffenbach, and A.S. Fauci, JAMA 321:451-452, 2019; DOI:10.1001/jama.2018.21167)
NEI: FAULTY MOLECULAR MASTER SWITCH MAY CONTRIBUTE TO RETINAL DEGENERATION
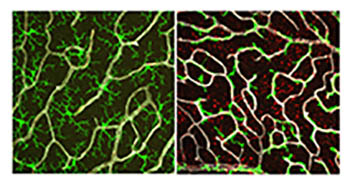
CREDIT: WENXIN MA, M.D., PH.D., AND WAI WONG, M.D., PH.D., NEI
In healthy retina (left), microglia (green) have a branched structure that covers the retina. Without TGF-beta signaling (right), microglia lose their branched structure and attach to blood vessels (white). Müller glia become abnormal and acquire activation markers (red).
In a recent study, NEI researchers explored the role of transforming growth factor–beta (TGF-beta) signaling in retinal health and disease. They found that a signaling pathway controlled by TGF-beta could be involved in the progression of age-related macular degeneration (AMD), a common cause of vision loss among older Americans. Interrupting TGF-beta signals to the microglia causes these cells to enter an activated, inflammatory state. These activated microglia damage the retina in a way that is similar to the cellular effects observed in AMD.
Using mouse models, the researchers ablated the TGF-beta receptor (TGFBR2) in the retinal microglia, which led to the loss of the microglia’s ability to sense TGF-beta signaling. Microglia in these mice had a stubbier morphology with less branching, which means less physical covering of the retina as well as disrupted microglia-synapse connection. The study also showed evidence of an activated, inflammatory gene-expression signature and of failure in the microglia’s ability to sense endogenous environmental signals. This homeostatic misbalance has cascading effects on the retina—the neighboring Müller cells, a type of retinal glial cell, began to show signs of distress, and retinal neurons began to fail and die.
Altogether this novel study has shown TGF-beta signaling in the retinal microglia is important for microglia homeostasis as well as retinal integrity. The pathological changes are similar to what happens in the progression to late AMD, indicating that microglia and TGF-beta signaling may help drive disease progression in humans. TGF-beta, therefore, may represent an important therapeutic target for treating AMD, according to the researchers. (NIH authors: W. Ma, S.M. Silverman, L. Zhao, R. Villasmil, M.M. Campos, J. Amaral, and W.T. Wong, eLife 2019; DOI:10.7554/eLife.42049)
NIDCD, NCCIH: GENETIC VULNERABILITY TO MENTHOL IN CIGARETTES
NIH researchers have found that a genetic variant is linked to the preference for menthol cigarettes among African-Americans. Menthol is a common cigarette flavoring that provides a minty taste and a cooling or soothing sensation in the mouth. Mentholated cigarettes have been gaining a larger share of the cigarette market in the United States even as overall smoking has declined over the past 50 years. There are also significant demographic differences in preference for menthol cigarettes. Nearly 83 percent of African-American smokers prefer mentholated cigarettes versus only 24 percent of white smokers and 32 percent of Hispanic smokers. In a collaborative effort, researchers from NIDCD, NCCIH, and other institutions set out to assess whether there is a genetic basis to this preference.
They analyzed protein-coding genes and took surveys of smoking habits in 1,300 adults (in groups that included multiethnic and African-American smokers), leading to the discovery of a genetic variant unique to African-Americans. A variant of the MRGPRX4 gene was found in five to eight percent of African-Americans and was absent from other demographics. Smokers with this variation were five to eight times as likely to reach for mentholated cigarettes as for nonmentholated ones.
The protein of MRGPRX4 was identified as a G-protein-coupled cell-surface receptor associated with detecting irritation rather than taste. Menthol suppresses the activity of this receptor, an effect that is enhanced in those with the variant. The study predicts that menthol provides an anesthetic-like effect through this signaling pathway, making mentholated cigarettes less irritating. While not the only factor in cigarette choice, a potential genetic vulnerability to mentholated cigarettes among African-Americans could be used to help inform public-health policy decisions about the use of menthol in cigarettes. (NIH authors: D. Risso, E. Sainz, A. Barik, C. Frigerio-Domingues, and D. Drayna, PLoS Genet 15:e1007916, 2019; DOI:10.1371/journal.pgen.1007916)
NCI: TREATING GLIOMA WITH AN NRF2 BLOCKADE
Glioma, a devastating tumor affecting thousands in the United States, is the most frequent malignant brain and central-nervous-system tumor, with an estimated 20,000 newly diagnosed cases each year. NCI researchers investigated a common type of IDH1-mutated glioma—and found that targeting its antioxidant pathway could be a successful treatment strategy. IDH1 mutations disrupt the reduction–oxidation reaction (redox) balance by promoting the production of reactive oxygen species (ROS), which are a natural byproduct of the normal metabolism of oxygen and have important roles in cell signaling and homeostasis.
The researchers used patient-derived glioma cells and mouse models to examine the metabolic signature and gene-expression profile in IDH1-mutated glioma. They also analyzed DNA oxidation and accumulation of ROS in tumor cells. They found that IDH1-mutated glioma cells exploit the function of nuclear factor (erythroid-derived 2)–like 2 (NRF2), which enables detoxification pathways to support the metabolic pathways and redox homeostasis in cancer cells. Using a preclinical xenograft model, the researchers showed that a pharmacologic blockade of NRF2 potently reduced tumor expansion, suggesting a novel therapeutic strategy for this type of malignancy. (NCI authors: Y. Liu, Y. Lu, O. Celiku, A. Li, Q. Wu, Y. Zhou, and C. Yang, J Natl Cancer Inst 111, 2019; DOI:10.1093/jnci/djy230)
NIEHS, NCI, NIDDK, NHLBI: ACTIVE GENES EXHIBIT VARIABLE INACTIVE PERIODS
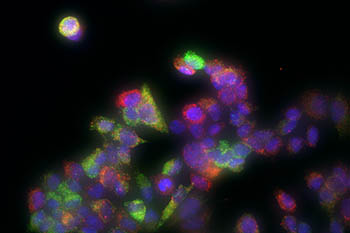
CREDIT: JOSEPH RODRIGUEZ, NIEHS; CARSON CHOW, NIDDK; AND DANIEL LARSON, NCI
Imaging of individual breast-cancer tumor cells shows two different estrogen-responsive genes, one highlighted in green and the other in red. Both genes show different levels of activity in different cells, although they were exposed to the same amount of estradiol.
New research calls into question the understanding of gene expression as an either/or proposition—genes are either on and active, or they are off and thus inactive. The findings indicate that active genes likely spend most of their time in an inactive phase called the deep repressive state. The results may lend insight into how variability in gene expression can contribute to disease. “Our study is one of the real firsts in observing how human genes respond to hormones in real time,” said lead study author Joseph Rodriguez, a new faculty member in NIEHS’s Epigenetics and Stem Cell Biology Laboratory. He led the project as a postdoctoral fellow in the lab of Dan Larson at NCI. These findings “add a new layer of regulation to the current model of transcription,” Rodriguez said.
Over the past decade, researchers have noticed variability in the way genes are turned on or off, even in identical cells in identical environments. One cell might be producing a couple dozen copies of a particular RNA message at the same time that another cell in that same tissue is churning out hundreds. This gene-expression noise gets worse with age and is a hallmark of cancer. “This noise exists in normal tissues,” said Rodriguez. “But in cancer, it appears to be hijacked to allow even more diversity in the tumor population, making it easier for tumor cells to develop resistance to treatment.”
Rodriguez used the CRISPR gene-editing technique to tag multiple alleles of the TFF1 gene in breast-cancer cells and then record in real time the response to estradiol by different alleles in the same cell. “We saw that some alleles were stuck in a long, deep repressive state that could last several days, even when other alleles nearby were flipping on and off,” Rodriguez said. “Human transcription is dynamic, more dynamic than we ever realized.”
Rodriguez has more questions about this deep repressive state that he hopes to answer in his new post at NIEHS. “Why does this happen? How does it change as a result of environmental exposure? How does it become dysregulated as we age?” he asked. Using his live-imaging technique, Rodriguez wants to look at how endocrine disruptors, which can mimic hormones such as estrogen, affect expression of genes like TFF1. (NIH authors: J. Rodriguez, G. Ren, C.R. Day, K. Zhao, C.C. Chow, D.R. Larson, Cell 176:213–226, 2019; DOI:10.1016/j.cell.2018.11.026)
This page was last updated on Tuesday, April 5, 2022
