Challenges to Training Artificial Intelligence with Medical Imaging Data
If you were going to train an artificial intelligence (AI) system to understand and accurately diagnose medical images, what kind of information do you think would be most effective: lots of general image data, or small amounts of specific data?
In my previous blog post, I briefly described our team’s work using data in the NIH Clinical Center’s picture archiving and communication systems (PACS) to build an AI system that can learn from radiology reports and images stored in PACS to generate descriptive keywords of new images independently of human eyes. Such a system could be used to provide first-impression summarizations of medical images and index large amounts of images so that doctors or researchers can search for images with certain characteristics, such as “images with brain tumor" or "magnetic resonance images of muscle.” However, our research objectives are even grander.
One of our main goals is to build an AI system that can make automatic disease diagnoses from patient scans, something that our proposed system is not quite able to do. To address its limitations, we used Metathesaurus (a medical semantic database of the Unified Medical Language System [UMLS]) and the RadLex radiology language database to detect disease-specific words used in radiology from the reports. In addition, we employed a negation detection algorithm to determine whether each detected disease word is mentioned positively (disease is present) or negatively (disease is absent) in the image descriptions. Then, we trained deep convolutional neural networks to detect the presence or absence of a disease in given medical images.
Below are two example outputs of the trained AI system, which generated keywords and detected the presence or absence of a disease with confidence levels. The original text from each image’s human doctor-generated radiology report is also shown on the right.
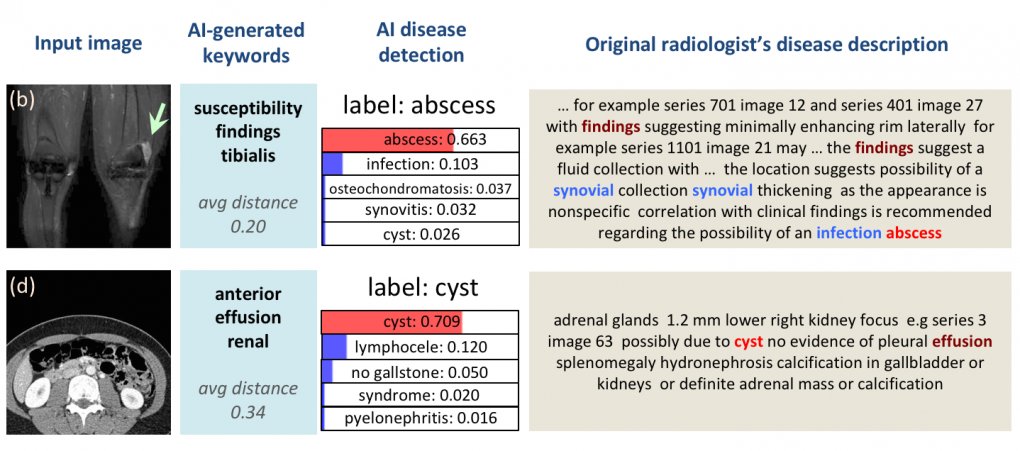
Example outputs for automated image interpretation, where the AI’s most probable disease detection matches the radiologist’s assigned label.
As we found, the AI system can detect the presence or absence of a disease in a given image, which supplements AI generated keywords towards the purpose of diagnosis. Still, two big challenges remain:
Challenge 1: Reports written by different doctors can vary widely, and multiple terminologies may be used to describe the same type of disease.
Matching variable descriptions to a single disease type and training an AI to understand them all is challenging. To train for more specificity in disease detection, we could only use about 10% of the entire data available in the PACS. By aiming to generate approximate keywords describing each image we could ignore description variability, but then we also had to be more specific in the text mining, which filtered out about 90% of the available data.
Challenge 2: Doctors sometimes write in an indefinite way when describing a disease.
"Possibility of an infection abscess," "not obviously cysts," or "possibly due to cyst" (as with image “d” above) are descriptors commonly used by doctors to advise further investigation. Without manually examining such images, it is challenging to derive any concrete findings from uncertain text descriptions. Indeed, it was hard to find any cyst in the “d” example image.
Nonetheless, having more data is always desirable when building a machine learning system, so that the system can better generalize beyond the examples it has seen in the training set. Our experience raises interesting questions regarding the trade-offs in designing an artificial intelligence to understand medical data: Do we go big and less specific with the data used for training, or small and more specific?
Look forward to my next post, where I will describe our efforts to train an AI system to understand medical images, using unrelated images of everyday objects.
More details about our study can be found in the published journal article: Shin et al., Interleaved Text/Image Deep Mining on a Large-Scale Radiology Database for Automated Image Interpretation, Journal of Machine Learning Research.
Related Blog Posts
This page was last updated on Wednesday, July 5, 2023
