Research Briefs
NIDDK, NHGRI, NCI, CC, NCBI: NIH STUDY FINDS GENETIC LINK FOR RARE INTESTINAL CANCER
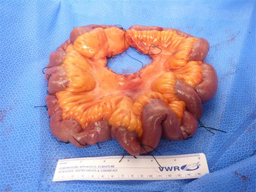
GASTROENTEROLOGY 2015
NIH researchers found a genetic link for small intestinal carcinoid, a rare digestive cancer. Shown is a section of the small intestine with the wide distribution of multiple carcinoid tumors identified by sutures.
Heredity accounts for up to 35 percent of small intestinal carcinoid, a rare digestive cancer, according to findings from a team of NIH researchers. Because the disease has long been considered to occur randomly rather than to be inherited, people with a family history are not typically screened. Conducted at the NIH Clinical Center, the study screened 181 people from 33 families, each with at least two cases of small intestinal carcinoid. The researchers discovered the disease in 23 people who had not yet developed symptoms and successfully removed all tumors in 21 of those people. Genetic linkage analysis revealed a target DNA region shared by all affected members of a particularly large family. Genome sequencing narrowed that finding to a gene defect passed from one generation to the next, suggesting that the gene is an inherited risk factor for the disease. (NIH authors: Y. Sei, S. Wank, S. Szymczak, Q. Li, J. Bailey-Wilson, et al., Gastroenterology DOI:10.1053/j.gastro.2015.04.008)
NIAID: NO EVIDENCE OF ACCELERATED EBOLA VIRUS EVOLUTION IN WEST AFRICA
The Ebola virus circulating in humans in West Africa is undergoing relatively few mutations, none of which appear to make the virus more severe or transmissible, according to a study by scientists at NIAID’s Rocky Mountain Laboratories (Hamilton, Montana). The study compares virus sequencing data from samples taken in 2014 from patients in Guinea, Sierra Leone, and Mali, and it finds that there appear to be no genetic changes that would increase the virulence or change the transmissibility of the circulating Ebola virus. Despite extensive human-to-human transmission during the outbreak, the virus is not mutating at a rate beyond what is expected. Further, based on the data, it is unlikely that the types of genetic changes thus far observed would impair diagnostic measures or affect the efficacy of candidate vaccines or potential virus-specific treatments. (NIAID authors: T. Hoenen, T. Schwan, R. Sakai, M. Niang, and M. Pineda, Science 348:117–119, 2015)
NIEHS: RESPONDING TO ENVIRONMENTAL CUES REQUIRES CROSS-TALK BETWEEN SIGNAL TRANSDUCTION AND TRANSCRIPTION
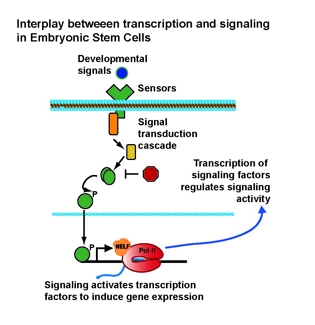
NIEHS researchers determined that control of developmental signaling in mouse embryonic stem cells (ESCs) can be accomplished through RNA polymerase II (Pol II) pausing, a phenomenon in which Pol II pauses during early transcription of messenger RNA. The finding is important because it sheds light on a mechanism that can be used to calibrate ESC responsiveness to environmental and developmental cues. Pausing of Pol II was known previously to be widespread in ESCs, but its function was unclear. A popular model predicted that disruption of pausing would cause genes involved in cell differentiation to be activated, resulting in spontaneous cell differentiation and development. The NIEHS team found, however, that the loss of pausing had the opposite effect: ESCs would be unable to respond to differentiation cues from their environment because developmental signaling machineries were repressed. Thus, mice lacking the key pause-inducing factor, negative elongation factor, died early in embryonic development because they were unable to detect and respond to information from their environment. (NIEHS authors: L.H. Williams, G. Fromm, N.G. Gokey, T. Henriques, G.M. Muse, A. Burkholder, D.C. Fargo, G. Hu, and K. Adelman, Mol Cell 58:311–322, 2015) [SUBMITTED BY ROBIN ARNETTE, NIEHS]
NCI: CIRCULATING TUMOR DNA IN BLOOD CAN PREDICT RECURRENCE OF LYMPHOMA
Measurement of circulating tumor DNA (ctDNA) in blood can be used to detect disease recurrence in patients with a curable form of cancer known as diffuse large B-cell lymphoma (DLBCL), the most common type of lymphoma. The disease recurs in up to 40 percent of patients and the recurrence is often incurable, particularly in those who progress early and/or have significant concentrations of tumor cells in their blood. NCI investigators found that measurement of ctDNA enabled the detection of microscopic disease before it could be seen on computerized tomography (CT) scans, which is the current standard for disease assessment. Monitoring for recurrence by testing blood samples may reduce the need for multiple CT scans that increase a patient’s exposure to radiation and add to health-care costs. Advances in the ability to monitor for disease recurrence earlier may also improve the ability of physicians to successfully treat the disease at the time recurrence is diagnosed.
The investigators used advanced sequencing techniques to analyze serum from 126 patients with DLBCL for the presence of ctDNA. All patients received therapy with the five drugs etoposide, prednisone, vincristine, cyclophosphamide, and doxorubicin, known as EPOCH, with or without the biologic agent rituximab, in clinical trials between May 1993 and December 2013. Serum samples were collected before treatment, during treatment, and for many years after therapy. The patients also had CT scans done at the same time as the blood testing as part of standard surveillance. They were followed for a median of 11 years after the completion of therapy.
Results of this study showed that, among the 107 patients who achieved complete remission, those who developed detectable ctDNA during surveillance were over 200 times as likely to have their disease progress as those who did not have detectable ctDNA. The researchers also found that measuring ctDNA enabled the detection of cancer recurrence a median of 3.4 months before clinical evidence of disease. In addition, the ctDNA test was able to predict which patients would not respond to therapy as early as their second cycle of treatment, a strategy known as interim monitoring. Interim ctDNA is therefore a promising biomarker to identify patients at high risk of not responding to treatment for their disease, according to the researchers. (NCI authors: M. Roschewski, K. Dunleavy, S. Pittaluga, M. Shovlin, E.S. Jaffe, L.M. Staudt, et al., Lancet Oncol DOI:10.1016/S1470-2045(15)70106-3)
NICHD: ANTI-HERPES DRUG MAY HELP CONTROL HIV
Valacyclovir, a drug commonly used to control the virus that causes genital herpes, appears to reduce the concentration of human immunodeficiency virus (HIV) in patients who do not have genital herpes, according to a study by researchers from NICHD, Case Western Reserve University (Cleveland), Emory University (Atlanta), and Asociación Civil Impacta Salud y Educación (Lima, Peru). The study enrolled 18 HIV-infected patients, none of whom were infected with herpes simplex virus–2 (HSV-2, the type that causes genital herpes), and treated them with valacyclovir. For 12 weeks, half of the enrolled patients took valacyclovir twice a day while the other half received a placebo. After two weeks, the placebo group received valacyclovir while the group originally treated with the drug switched to the placebo.
When the patients took valacyclovir, their blood HIV concentrations declined significantly. The researchers conducted a genetic analysis and found that the HIV in the study volunteers did not develop resistance to valacyclovir. But because HIV has a history of becoming resistant to the drugs used to treat it, the researchers do not discount the possibility that the virus could develop resistance to valacyclovir with longer treatment. If valacyclovir’s effectiveness against HIV can be confirmed in a larger cohort, it could be added to the mix of drugs used to suppress the virus and might prove especially helpful in cases in which HIV has developed resistance to other drugs. (NICHD authors: C. Vanpouille, A. Lisco, J. Grivel, and L. Margolis, Clin Infect Dis DOI:10.1093/cid/civ172)
NCATS, NIDDK: ALLERGY DRUG INHIBITS HEPATITIS C IN MICE
NCATS and NIDDK researchers have found that an over-the-counter drug used to treat allergy symptoms limited hepatitis C virus (HCV) activity in infected mice. Using a high-throughput screening process, the researchers identified chlorcyclizine HCl (CCZ) as a potent inhibitor of hepatitis C. HCV causes liver inflammation and often leads to serious complications such as cirrhosis. Early diagnosis and treatment of HCV can prevent liver damage. Drugs are available to treat HCV, but costs can reach tens of thousands of dollars. The study found that CCZ blocked the early stage of HCV infection, likely by impairing the ability of the virus to enter human liver cells grafted in the mice. The outcome was similar to that of commonly used antiviral drugs but without those drugs’ toxic side effects. The researchers will next study how the drug affects people. CCZ is currently used for the treatment of allergies, not for HCV. The scientists caution that people should not take CCZ to treat their hepatitis C until it has been demonstrated that CCZ can be used safely and effectively for that purpose. (NIDDK authors: S. He, B. Lin, T.J. Liang, et al.; NCATS authors: X. Hu, J. Xiao, et al., Sci Transl Med DOI:10.1126/scitranslmed.3010286)
NIDCR: SIGNALS FROM EPITHELIAL PROGENITOR CELLS PROMOTE ORGAN INNERVATION DURING DEVELOPMENT
A key process in embryonic development is the integration of different cell types to form organs. In some developing organs such as the salivary glands and pancreas, innervation occurs during organ formation, with neurons clustering together and sending out axonal projections along the epithelium in three dimensions. Using ex vivo tissue cultures and molecular techniques, scientists in the National Institute of Dental and Craniofacial Research (NIDCR) have identified cells and signals that drive innervation within the embryonic salivary gland. They are the first to describe how the salivary gland regulates its own innervation.
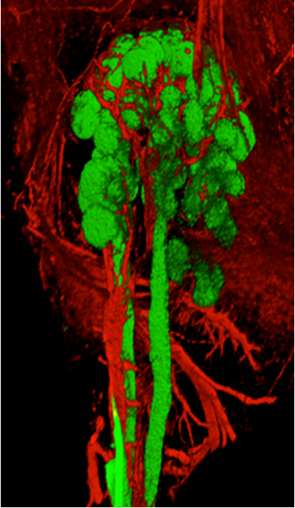
WENDY M. KNOSP, NIDCR
Scientists in NIDCR have identified cells and signals that drive innervation within the embryonic salivary gland. Shown: image of an immunostained salivary gland (green, grapelike clusters) and the surrounding nerves (red).
In a mouse embryo, the salivary glands bud, branch, and form hollow ducts within two weeks of conception. The resulting architecture, which resembles a cluster of grapes, depends on a precise coordination of molecular signals from epithelial cells, mesenchymal cells, nerve cells, and other cell types. NIDCR’s developmental cell biologists in the Matrix and Morphogenesis Section have identified the signals that initiate the development of the innervation of the salivary gland. Senior investigator Matthew P. Hoffman and colleagues recently reported in Developmental Cell that Wnt signals from keratin-5-positive (K5+) progenitor cells in each developing salivary gland’s epithelial compartment are driving the process of innervation.
Using double-knockout mouse embryos that had been genetically modified to lose the ability to properly regulate fibroblast growth factor (FGF) signaling, they discovered a striking loss of innervation in the developing salivary glands. They also found a dramatic reduction in epithelial Wnt expression in the knockout salivary glands and demonstrated that FGF signaling suppresses Wnt expression. In addition, they found that Wnt signaling is essential for nerve-cell survival and proliferation and thus for innervation of the gland.
To rescue innervation in the knockout salivary glands, the scientists administered an oral formulation of a Wnt activator called CT99021 while reducing FGF signaling. In addition, the NIDCR researchers were able to extend the findings to the embryonic pancreas.
Their work builds on their previous finding published in Science in 2010 that the nervous system maintains the K5+ cells in the epithelial compartment; without the appropriate signals from nerves, the K5+ cells differentiate, and gland development is impaired (Science 329:1645–1647, 2010). These studies are part of the groundwork for future clinical application of tissue repair and regeneration by re-establishing communication between nerves and transplanted progenitor cells. (NIDCR authors: W.M. Knosp, S.M. Knox, I.M.A. Lombaert, C.L. Haddox, V.N. Patel, and M.P. Hoffman, Dev Cell 32:667–677, 2015)
This page was last updated on Monday, April 25, 2022
