New Methods
Laser Capture Micro-Dissection Gets Automated
Michael A. Tangrea, NCI
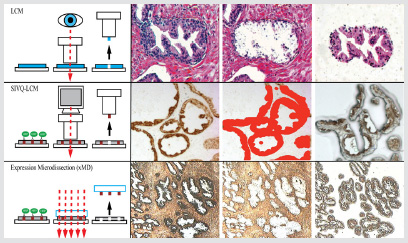
NIH researchers use three different types of laser-capture microdissection (LCM) technologies to dissect tissue such as the human prostate gland shown here. From top, the traditional LCM and the more automated systems—spatially invariant vector quantization (SIVQ)–LCM and expression microdissection (xMD).
You’d think it would be hard to top laser-capture microdissection, or LCM. This technique, developed at the NIH in the 1990s, uses lasers and other specialized instruments to carve out sections of tissue as small as a few cells from across a complex specimen and then lift them away almost magically, undamaged, for further analysis.
For the operator, it’s like playing a video game, zapping and lifting with trigger buttons, guided by microscopic visualization.
LCM instruments are in pathology labs across the globe and have generated close to 3,000 scientific publications and hundreds of millions of dollars in sales, licensed through Arcturus Engineering/Life Technologies and other companies. The inventors include NCI’s Michael Emmert-Buck, NICHD physicist Robert Bonner, CIT engineer Tom Pohida, and Lance Liotta, former deputy director for intramural research.
But manual laser slicing is so old school. Now, NIHers are further automating the process, developing parallel technologies that employ immunohistochemistry and image-analysis software to make the microdissection process nearly operator-independent.
Their goal, said Emmert-Buck, is to bring laser dissection to the clinic to help precipitate the era of personalized medicine. The new techniques are ideal for studies requiring large amounts of material, such as proteomics and genomic assays. For example, to understand how a drug will interact with a specific patient’s tumor, one often needs thousands of precisely dissected cells. Automated methods can dissect cells much faster than the manual-based traditional LCM.
“They are trying to replace me,” laments Jaime Rodriguez-Canales, a molecular pathologist in the NCI Laboratory of Pathology’s LCM Core, which has assisted with dissections for countless NIH labs over the past decade.
Although sounding like a modern-day John Henry—the American folk hero who proved he could hammer rocks faster than a steam-powered hammer—Rodriguez-Canales is being facetious. Yes, it is true that his steam-hammer-wielding colleague, biomedical engineer Jeffrey Hanson, was slicing out dozens of cancer cells. But expert molecular pathologists will be needed more than ever to prep specimens for dissection and then discern the meaning of the data that comes out.
One new dissection method is expression microdissection (xMD), which uses a targeting probe for cell procurement in place of the operator-based, cell-by-cell LCM selection process. This technology recently was adapted for commercially available LCMs, enabling investigators outside of the NIH to use it. The xMD method might allow subcellular dissections, a notable advance for the field.
The other new method is spatially invariant vector quantization (SIVQ)-LCM, an image-based dissection technology developed by the NIHers and University of Michigan School of Medicine (Ann Arbor, Mich.). SIVQ uses mathematical modeling and image processing to locate phenotypically similar cells across entire tissue sections.
The complementary LCM, xMD, and SIVQ-LCM technologies can be applied widely in research and clinical settings. For example, a pathologist analyzing a patient biopsy to provide a standard histopathological diagnosis can select a molecular target panel to be studied using the optimal technique—SIVQ–LCM, xMD, or standard LCM—for harvesting cells.
The pathologist then can provide information to clinicians on clinical diagnosis and status of biological markers, useful for determining patient prognosis and selecting the most efficacious treatment. The NIH Clinical Center is evaluating such a workflow.
The NIH intramural program is an ideal incubator for these technologies because of the confluence of clinicians, engineers, pathologists, and molecular biologists, said Emmert-Buck. Although both xMD and SIVQ–LCM are still in the early commercialization stage, he welcomes NIH collaborators to beta-test the new methods.
As for the good-natured joking about John Henry, Emmert-Buck reassured Rodriguez-Canales: “A good pathologist with a good tool is just going to be better,” he said. “The history of science shows that new technologies lead to new discoveries that benefit everyone.”
This page was last updated on Monday, May 2, 2022
