Mapping Translocations
New Technique Identifies First Events in Tumor Development
NIGMS
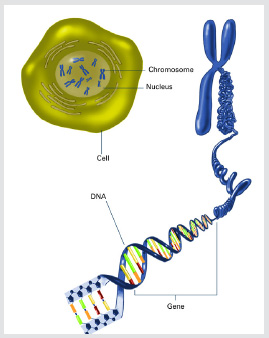
The long, stringy DNA that makes up genes is spooled within chromosomes inside the nucleus of a cell. (Note that a gene would actually be a much longer stretch of DNA than what is shown here.)
NHGRI
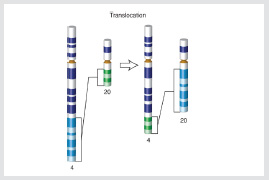
Translocations occur when a broken strand of DNA from one chromosome is erroneously joined with that of another chromosome. Translocations can result in tumors.
A novel technique that enables scientists to measure and document tumor-inducing changes in DNA is providing new insights into the earliest events involved in the formation of leukemias, lymphomas, and sarcomas and could potentially lead to the discovery of ways to stop those events.
A team of researchers at NIAMS, NCI, and Rockefeller University (New York) developed a high-throughput technique—called translocation capture sequencing (TC-Seq)—to document chromosomal rearrangements, or translocations, in primary cells. They reported their findings recently in the journal Cell (Cell DOI 10.1016/j.cell.2011.07.048).
Translocations occur when a broken strand of DNA from one chromosome is erroneously joined with that of another. Such irregularities can be beneficial—they may enable the immune system to respond to a vast number of microorganisms and viruses—but they can result in tumors. Translocations can take place during the course of normal cell division when each chromosome is copied verbatim to provide genetic information for the daughter cells.
“The cell expresses specific enzymes whose primary purpose is to repair such lesions effectively, but when the enzymes mistakenly join pieces of two different chromosomes, the cell’s genetic information is changed,” said Rafael Casellas, senior investigator in the NIAMS’s Genomics and Immunity Section.
Casellas likens the phenomenon to breaking two sentences and then rejoining them incorrectly. For example, “The boy completed his homework” and “The dog went to the vet” might become “The dog completed his homework” and “The boy went to the vet.” When a cell gets nonsensical information such as this, it can become deregulated and even malignant.
Scientists have known since the 1960s that recurrent translocations play a critical role in cancer. What was unclear was how these genetic abnormalities are created, since very few of them were studied, and only within the context of tumors, said Casellas. To better understand the nature of these tumor-inducing rearrangements, the authors created a system to visualize their appearance in normal, nontransformed cells.
Using the TC-Seq system they created, the scientists investigated how oncogenic rearrangements occur. First, they introduced enzymes that recognize and cause damage at a particular sequence in the DNA into cells from mice, thereby constructing a genome in which a unique site is broken continuously.
Next, they used polymerase chain reaction—a technique to quickly amplify short sequences of DNA—to check all of the sites in the genome that would get translocated to this particular break. The researchers examined more than 180,000 chromosomal rearrangements from 400 million B cells.
Based on this large data set, the scientists made several important observations about the translocation process. They learned that most of the translocations involve gene domains rather than the space on the DNA between the genes.
They also found that most translocations target active genes, with a clear bias for the beginning of the gene, as opposed to its middle or end. The team also showed that a particular enzyme that normally creates DNA breaks in B cells dramatically increases the incidence of translocations during the immune response.
This feature explains the long-standing observation that more than 95 percent of human lymphomas and leukemias are of B-cell origin.
This molecular understanding of translocation hot spots “is allowing us to understand how tumors are initiated,” said Casellas. “It is the kind of information that in the near future, might help us prevent the development of cancer.”
This page was last updated on Monday, May 2, 2022
