Demystifying Medicine Explores the Origin of Life
Evolutionary Biologist and Cell Biologist Share Their Views

The Demystifying Medicine presenters answered questions from the audience about their views on the origin of life. Clockwise from top left: Win Arias (host and founder of the Demystifying Medicine course), chemist Nick Lane, and cell biologist Jennifer Lippincott-Schwartz.
Four billion years ago, life arose on Earth from nonliving matter. About two billion years later, more complex and multicellular life forms began to emerge. Scientists continually debate how life began, but two of them— evolutionary biologist Nick Lane and cellular biologist Jennifer Lippincott-Schwartz—shared their theories with an audience that had gathered (online) for a Demystifying Medicine seminar on “The Origin of Life.”
Energy flow shaped evolution
Nick Lane, who is a professor of evolutionary biochemistry at University College London (London), talked about how energy flow has shaped evolution. The focus of his research is on chemiosmosis, the process of moving ions across biological membranes and generating an electrochemical gradient, which in turn drives the synthesis of adenosine triphosphate (ATP). ATP is the energy-carrying molecule found in the cells of all living organisms.
He explained how hydrothermal vents on the ocean floor may hold some clues to the origin of life. NASA geochemist Michael Russell proposed that the first life forms started in these vents, which contained hydrogen and methane gases. The vents’ pores resemble the outer membranes of cells. A proton gradient forms on the membranes and drives carbon dioxide (CO2) fixation, the process by which plants and algae create energy and food from CO2 in the air. To recreate this reaction, Lane created an origin-of-life reactor (protocell) in the lab. He found that in alkaline hydrothermal conditions, protons could cross the artificial barrier and that protocells with bilayer membranes generate spontaneously. An aqueous environment favors the formation of ATP.
Origin of eukaryotic cells
CREDIT: JENNIFER LIPPINCOTT-SCHWARTZ
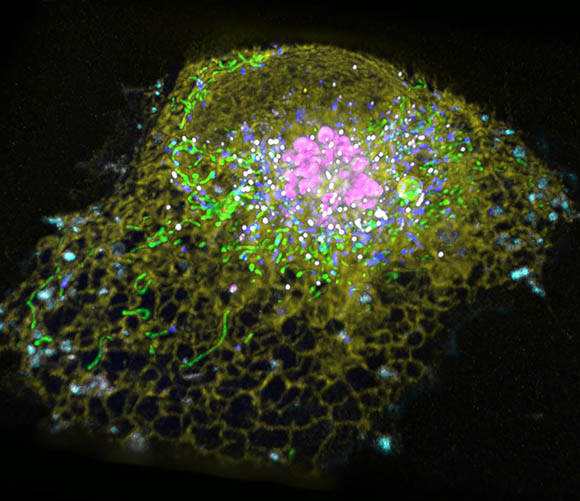
Jennifer-Lippincott-Schwartz uses a variety of advanced imaging techniques to better understand how cells function in different organs. Shown: Hyperspectral imaging that shows different color signals for six types of membrane-bound organelles in an eukaryotic cell: endoplasmic reticulum, Golgi, lysosome, lipid droplet, peroxisome, and mitochondria.
In the next presentation, Jennifer Lippincott-Schwartz, a former NIH investigator and now a senior group leader at Howard Hughes Medical Institute’s Janelia Research Campus (Ashburn, Virginia), described her studies on the origin of eukaryotic cells.
When life began to emerge, prokaryotes (single-cell organisms) slowly evolved into eukaryotes (multicellular organisms with a nucleus and other membrane-bound organelles). Lippincott-Schwartz talked about two models of eukaryotic origin: stepwise evolution in which prokaryotic membranes became more complex and evolved into proto-eukaryotes and then eukaryotes; metabolic symbiosis in which two prokaryotic cells interact with each other by providing hydrogen, CO2, gene transfer, and energy. In the latter interaction, archaea (primitive prokaryotes that lack a nucleus) were the host cells and bacteria acted as mitochondria; the pairing resulted in eukaryotic cells.
One difference between prokaryotic and eukaryotic cell membranes is that prokaryotes lack cholesterol and other lipids. Lippincott-Schwartz proposed that the appearance of cholesterol and sphingolipids in the eukaryotic membrane was a turning point in eukaryotic cell evolution. Cholesterol and sphingolipids give membranes the ability to phase partition protein components, especially at the cell surface and in Golgi membranes. Protein sorting within membranes, in turn, enables more elaborate complex structures, including membrane trafficking pathways and distinct organelles, such as the lysosome, endosome, endoplasmic reticulum, and Golgi apparatus.
Lippincott-Schwartz told the audience that to confirm these theories, she would need to conduct more elaborate experiments and develop mathematical models to study cellular pathways.
To watch a videocast of this seminar, presented on March 1, go to https://videocast.nih.gov/watch=44357. For more about the course—which is administered by the NIH Clinical Center—including background information about each topic, speakers’ CVs, selected references, and PowerPoint presentations, go to https://demystifyingmedicine.od.nih.gov. The Demystifying Medicine course, established and run by Irwin Arias, is in its 21st year and aims to bridge exciting developments in medicine with advances in the basic biological and engineering sciences. When possible, a live patient introduces the course topic by discussing their disease, followed by a physician–scientist who describes clinical aspects of the disease including therapy and challenges, followed by a bench scientist who describes what we know and don't know regarding mechanisms. The goal is to challenge interest and communication between attendees and speakers. You can view past videocasts (dating back to 2003) at https://videocast.nih.gov/PastEvents.asp?c=45.

Devika Bose is a molecular biologist and a laboratory technician in the National Eye Institute. She works in Kapil Bharti’s lab in the Ocular and Stem Cell Translational Research Section, and uses stem-cell-derived retinal pigment epithelial cells to study different retinal degenerative diseases. Outside of work, she enjoys traveling, cooking, hiking, binge watching programs on Amazon Prime, and reading.
This page was last updated on Tuesday, May 17, 2022
