News You Can Use
New Web Application to Study Cellular Responses to Glucocorticoids
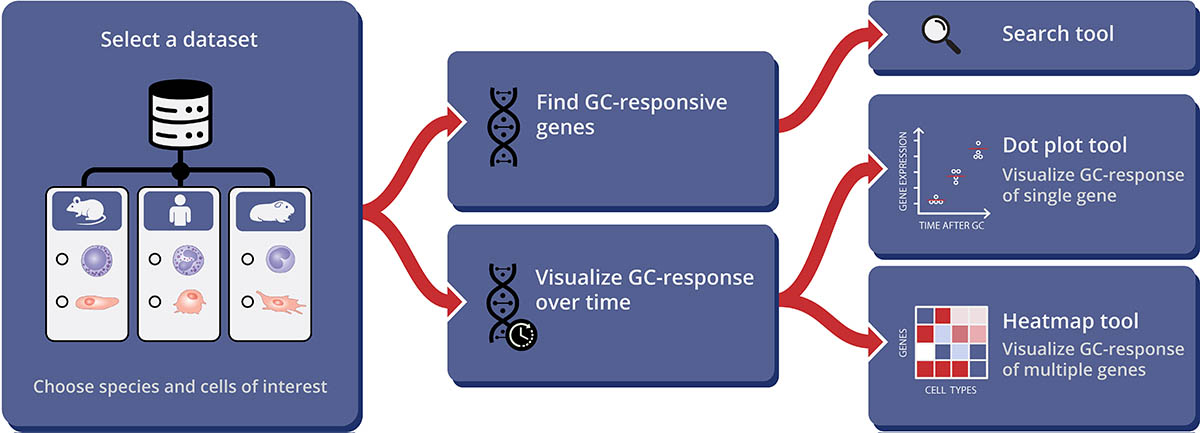
CREDIT: MARTYN GREEN, NIAMS
Typical workflow for using the GCgx tool. The user selects a dataset containing cells of interest for a chosen species then searches for GC-responsive genes or visualizes GC-response over time results as a dot plot (for a single gene) or a heatmap (for multiple genes).
Glucocorticoids are a powerful class of steroid drugs used for anti-inflammatory and immunosuppressive therapy and to fight an overactive immune system. During the COVID-19 pandemic, for example, dexamethasone and other glucocorticoids were successfully used to treat patients with severe COVID-19 by mitigating the systemic inflammatory response that can lead to lung injury and multisystem organ dysfunction. But these drugs can also have serious side effects that affect every organ system. Although the drugs have been around for more than 70 years, scientists have a poor understanding of how glucocorticoids regulate the immune system and the mechanisms by which they cause toxicity in different organs. Many important experiments have been performed in cell-culture or animal models. However, recent studies have revealed that each type of cell responds very differently to glucocorticoids, so those findings can’t be easily extrapolated to determine how human cells might react.
Now there’s a new web tool that will allow scientists to study how different human cell types will respond to glucocorticoids. The tool, called GCgx, was developed by Luis Franco’s lab at the National Institute of Arthritis and Musculoskeletal and Skin Diseases (NIAMS) and scientists at Bioinformatics and Computational Biosciences Branch at the National Institute of Allergy and Infectious Diseases (NIAID). Franco is an Earl Stadtman Investigator and an NIH Distinguished Scholar, and has a secondary appointment at NIAID. The tool is described in the Journal of Molecular Endocrinology (J Mol Endocrinol 68:B1–B4, 2022).
“GCgx is a scientific web application that allows investigators to quickly find answers to questions like, ‘Are my genes of interest responsive to glucocorticoids in specific cell types?’” said Franco. “If so, how does their level of expression change over time, and how statistically significant are the differences?”
GCgx is a mobile-friendly tool that can quickly display information about the transcriptional response to glucocorticoids from individual cell types. The tool has an extensive dataset that was generated in Franco’s lab and is based on total RNA sequencing in nine primary human cell types: B cells, CD4+ T cells, endothelial cells, fibroblasts, monocytes, myoblasts, neutrophils, osteoblasts, and preadipocytes. New datasets will be added over time, based on user input.
“GCgx will make it much easier to share data in a ready-to-query format with the rest of the research community, hopefully enabling better-informed experiments,” said Franco.
Researchers can use GCgx’s GC-responsive genes search function to determine contrasting gene responses in immune cells versus non-immune cells. GCgx’s heatmap function can simultaneously visualize the response to glucocorticoids across many genes and cell types within a given dataset. In addition, the dot plot function can be used to display the transcript abundance of a single gene of interest before and after glucocorticoid treatment.
“I am glad that Luis and colleagues developed the GCgx tool, because the utility of GCgx will be appreciated broadly by many groups,” said Earl Stadtman Investigator Mia Sung from the National Institute on Aging. Using published data from Franco’s lab and other groups, GCgx provides “a quick and easy way to query gene regulation of [glucocorticoids] in many human and mouse cell types. The GCgx team seems ready to improve the tool further with feedback and data contributions from users.”
Franco came up with the idea for GCgx in early 2020, when much of NIH’s bench research suddenly came to a halt due to the COVID-19 pandemic. He wanted to come up with a project that could be done outside of the lab and that could be accomplished using available data. Qilin Cao, postbaccalaureate fellow and the lead author of the aforementioned paper, took the idea and developed the web application.
“I think this is an example of how NIH postbacs can really make lasting contributions to their field of work,” said Franco.
Franco hopes that a better understanding of how glucocorticoids work will enable scientists to develop effective treatments with fewer side effects. GCgx is free, secure, and available to intramural researchers as well as those outside of the NIH. For more information on how to use GCqx, go to https://gcgx.niaid.nih.gov.

Satabdi Nandi, a postdoctoral fellow in the Laboratory of Molecular Biology and Immunology in the National Institute on Aging, is investigating the generation of antibody diversity in mouse B cells. Outside of work, she enjoys visiting museums to learn about the past, diverse cultures, and the motivation behind artists’ creations.
This page was last updated on Tuesday, May 17, 2022
