Research Briefs
NIDCR, NIAID, NIDCD, NCI: IMMUNE CULPRITS LINKED TO INFLAMMATION AND BONE LOSS IN GUM DISEASE
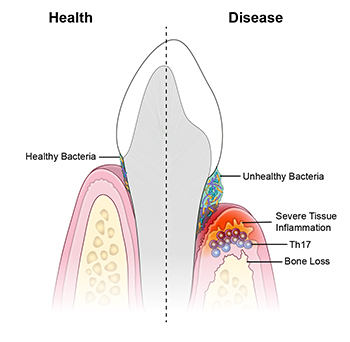
CREDIT: NIDCR
NIDCR: A new study suggests that periodontal disease is driven by Th17 immune cells, which are triggered by an unhealthy bacterial community.
An unhealthy population of microbes in the mouth triggers specialized immune cells that inflame and destroy tissues, leading to periodontal disease, according to a study led by researchers from NIDCR and the University of Pennsylvania School of Dental Medicine (Philadelphia). Periodontal disease affects nearly half of American adults over age 30 and 70 percent of adults 65 and older. The researchers observed that T helper–17 (Th17) cells were much more prevalent in the gum tissue of people with periodontitis than in the gums of their healthy counterparts, and that the amount of Th17 cells correlated with disease severity.
Th17 cells generally live in barrier sites—such as the mouth, skin, and digestive tract—and are known to protect against oral thrush, a fungal infection of the mouth. But they are also linked to inflammatory diseases such as psoriasis and colitis. The scientists found that, in mice with periodontitis, eliminating oral microbes prevented the expansion of Th17 cells in the gums while leaving other immune cells unaffected, suggesting an unhealthy bacterial population triggers the accumulation of Th17 cells. Blocking Th17 cells reduced the expression of genes involved in inflammation, tissue destruction, and bone loss, suggesting that Th17 cells may mediate these processes in periodontitis.
The researchers also studied a group of 35 patients, at the NIH Clinical Center, with a gene defect causing them to lack Th17 cells. The patients were less susceptible to gum disease and had less inflammation and bone loss compared with age- and gender-matched volunteers. The findings could have implications for new treatment approaches for gum disease and periodontitis. (NIH authors: N. Dutzan, L. Abusleme, T. Greenwell-Wild, C.E. Zuazo, T. Ikeuchi, L. Brenchley, C. Hurabielle, D. Martin, R.J. Morell, A.F. Freeman, V. Lazarevic, G. Trinchieri, S.M. Holland, Y. Belkaid, and N.M. Moutsopoulos, Sci Trans Med 10:eaat0797, 2018; DOI:10.1126/scitranslmed.aat0797)
NIEHS: GENETICS AND POLLUTION DRIVE SEVERITY OF ASTHMA SYMPTOMS
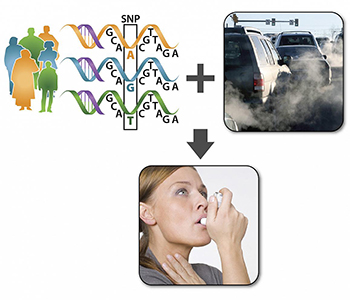
CREDIT: NIEHS
The research suggests when individuals with specific variations in certain genes are exposed to traffic pollution, they display more intense asthma symptoms than people that lack those same gene variations.
Some asthma sufferers have more severe reactions than others to air pollution caused by traffic. NIEHS researchers and others may have figured out why: It has to do with different genetic profiles. The scientists examined four particular single-nucleotide polymorphisms (SNPs) involved in a biochemical pathway that leads to inflammatory responses. They gathered information about the SNPs, severity of asthma symptoms, and residential addresses of 2,704 participants with asthma. Using the SNPs data, they divided the participants into three groups: hyper-responders, or those very sensitive to air pollution and likely to develop inflammation; hyporesponders, or those insensitive to air pollution and less likely to develop inflammation; and those in-between. With the help of collaborators at Rice University (Houston), the team used the participants’ addresses to calculate their distance from a major road.
The researchers found that asthma sufferers who were hyper-responders and lived closer to heavily traveled roads had the worst asthma symptoms, such as difficulty breathing, chest pain, cough, and wheezing, compared with the other groups. In contrast, asthma patients who were hyporesponders and lived further away from busy roads had milder symptoms. “Based on this research, we could propose that hyper-responders who are exposed to traffic pollution receive air-purification intervention, such as HEPA [high-efficiency particulate air] filters, for their home[s],” said Stavros Garantziotis, medical director of the NIEHS Clinical Research Unit and one of the authors of the study. (NIH authors: S.H. Schurman, M.A. Bravo, C.L. Innes, W.B. Jackson 2nd, J.A. McGrath, M.L. Miranda, and S. Garantziotis, Sci Rep 8:Article number 12713, 2018; DOI:10.1038/s41598-018-30865-0)
NIAID: PROBIOTIC BACILLUS ELIMINATES STAPHYLOCOCCUS BACTERIA

CREDIT: NIAID
Woman selling vegetable snacks in a Thai market—a possible source of probiotic Bacillus spores.
A new method of battling antibiotic-resistant superbugs is on the horizon. Researchers at the NIAID and two universities in Thailand revealed how oral Bacillus, a bacterium found in probiotic digestive supplements, can be used to fight superbugs such as methicillin-resistant Staphylococcus aureus (MRSA). MRSA can cause widespread infection of the skin, bloodstream, and lungs. However, when undisturbed, S. aureus lives benignly in the gut and nose of 30 percent of the general population. It only causes serious illness when the immune system is compromised or the bacteria break into the bloodstream through an exposed wound.
The typical treatment entails the administration of antibiotics, which are not always effective and may even result in the S. aureus becoming resistant to them. NIH researchers and their colleagues found that in 200 volunteers from rural Thailand (who were not expected to be as affected by food sterilization or antibiotics as people in urban areas), no S. aureus were found in gut and nose samples when Bacillus were present.
Then using chromatography and mass spectrometry techniques, the researchers discovered a class of lipopeptides (molecules that are part peptide and part lipid) called fengycins that inhibit S. aureus growth. The researchers showed that in mice, the fengycins eliminate S. aureus by inhibiting S. aureus quorum sensing—a process through which bacteria respond to their population density by altering gene regulation.
The next step is to test probiotics containing Bacillus subtilis to see whether they can eliminate S. aureus in people. (NIH authors: P. Piewngam, Y.Zheng, T.H. Nguyen, S.W. Dickey, H.-S. Joo, A.E. Villaruz, K.A. Glose, E.L. Fisher, R.L. Hunt, B. Li, J. Chiou, G.Y.C. Cheung, and M.L. Otto, Nature 2018; DOI:10.1038/s41586-018-0616-y)
NCCIH, NINDS: BROADER UNDERSTANDING OF HIGH-IMPACT CHRONIC PAIN
NIH researchers with colleagues at Kaiser Permanente Washington Health Research Institute (Seattle) have come up with a new classification for a type of chronic pain that leaves its sufferers so disabled that they can’t work and/or have trouble with routine self-care activities.
Of the 40 million people in the United States who have chronic pain, almost a quarter have what the researchers are calling high-impact chronic pain (HICP)—pain lasting three or more months in conjunction with at least one major activity restriction. HICP is commonly confused with chronic pain, which is defined by the duration of pain and not its progression to disability.
Using data from the 2011 National Health Interview Survey (15,670 adults), the researchers assessed the prevalence, characteristics, health status, and health-care usage of the chronic-pain population. About 4.8 percent of the country’s population experience HICP; 83 percent of people with HICP were unable to work and one-third had difficulty with self-care activities. The people with HICP experienced higher levels of anxiety, cognitive difficulty, and health-care usage. By differentiating the different types of chronic pain, the researchers hope to refine clinical treatments and identify interventions that can shape policy and improve the health of those with HICP. (NIH authors: M.H. Pitcher, M. Von Korff, M.C. Bushnell, and L. Porter, J Pain 2018; DOI:10.1016/j.jpain.2018.07.006)
[BY AUTUMN HULLINGS, NCI]
NICHD, NIMH: NEURONS ABSORB AND RELEASE WATER WHEN FIRING
Conventional functional magnetic resonance imaging (fMRI) technologies track neuronal activity indirectly by recording changes in blood flow and blood oxygen concentration as a proxy for metabolic activity. But researchers from NICHD and NIMH, and collaborators from other universities may have found a better way to measure brain activity: tracking active water cycling (AWC), which, they found, is characteristic of neuronal firing. The scientists made the discovery by simultaneously tracking neuronal activity and transmembrane water cycling in cultures of rat neurons. When neurons communicate, potassium and sodium ions move through the cell membrane. When the rat neurons were stimulated to fire, fMRI imaging revealed that the ion exchanges were accompanied by an increase in the number of water molecules moving into and out of the cell. This novel characteristic could lead to the development of an improved method of monitoring the brain’s circuitry by tracking AWC using fMRI. Although only proven in cell culture so far, this method could lead to a better understanding of the brain’s electrochemical activity and function, and ultimately of neural disease. (NIH authors: D. Plenz and P.J. Basser, Magnetic Resonance in Medicine, 2018; DOI:10.1002/mrm.27473)
NIAID: SCIENTISTS DEVELOP NOVEL VACCINE FOR LASSA FEVER AND RABIES
Scientists at NIAID and three universities have designed and tested a dual vaccine to protect people from both rabies and Lassa Fever (LF), which is highly fatal and for which no vaccine exists. LF can have varying symptoms in humans, ranging from mild illness to hemorrhage, and is widespread in West African countries such as Nigeria—which this year experienced its highest incidence of the disease. Africa is also at high risk for rabies in humans. The investigational vaccine, called LASSARAB, is an inactivated recombinant vaccine that uses a weakened rabies-virus vector and genetic material from the Lassa virus. It showed promise in preclinical testing in mice and guinea pigs. This multi-institutional effort represents an important preliminary advancement in providing a vaccine against LF. The researchers plan to evaluate the vaccine in nonhuman primates before conducting clinical trials. (NIH authors: K.R. Hagen, K. Cooper, P.B. Jahrling, and R.F. Johnson, Nature Commun 9:4223, 2018; DOI: 10.1038/s41467-018-06741-w)
NICHD, NCI, NIDDK, CC, NHLBI: INTERRUPTING SEDENTARY BEHAVIOR TO REDUCE METABOLIC RISK IN CHILDREN
Sedentary children are at greater risk of developing abnormalities in glucose homeostasis (the way blood sugar is regulated)—and at risk for developing diabetes—especially if they are overweight or obese.
It’s well known that supervised exercise improves glucose homeostasis. The trouble is, once the supervised training ends, the good effects disappear. In an effort to come up with alternative approaches, NICHD researchers and others sought to determine whether brief interruptions in sedentary behavior—such as short bouts of walking—would improve glucose metabolism. They designed a study in which 35 children ages 7–11 years who were overweight or obese underwent two experimental conditions. One group did prolonged sitting for three hours; the other group did interrupted sitting (three minutes of moderate-intensity walking every 30 minutes for three hours). Insulin, C-peptide, and glucose were measured every 30 minutes during an oral glucose-tolerance test.
The researchers concluded that interrupting sitting with brief moderate-intensity walking improved glucose metabolism. Interrupting sedentary behavior may be a promising intervention strategy for reducing metabolic risk in such children. “If this intervention provides sustained improvement in glucose metabolism . . . widespread implementation into school or after-school care centers could provide notable improvement in glucose homeostasis in the community setting and potentially slow the onset of type 2 diabetes in youth,” the authors wrote in their paper. (NIH authors: M.M. Broadney, B.R. Belcher, D.A. Berrigan, R.J. Brychta, I.L. Tigner Jr., F. Shareef, A. Papachristopoulou, J.D. Hattenbach, E.K. Davis, S.M. Brady, S.B. Bernstein, A.B. Courville, B.E. Drinkard, K.P. Smith, D.R. Rosing, P.L. Wolters, K.Y. Chen, and J.A. Yanovski, Diabetes Care 41:2220–2228, 2018; DOI:10.2337/dc18-0774)
NIDA: BRAIN ENSEMBLES THAT TUNE ON OR OFF TO SOCIAL EXPLORATION
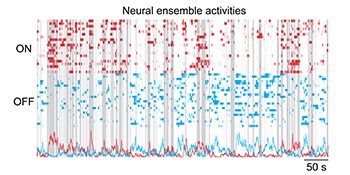
CREDIT: BO LIANG, NIDA
NIDA: The above plots show the ON (red) and OFF (blue) neural-ensemble activities recorded by a head-mounted miniature microscope when a mouse freely interacted socially with another mouse. Gray bars represent the period when the mouse explored his environment including possible social interactions.
Difficulties with social behavior can reflect a variety of psychiatric conditions, including schizophrenia and autism spectrum disorders. Some chronic substance-use behaviors could be attempts to minimize these social deficits. Knowing more about how the brain functions in social situations is critical to finding solutions to a broad range of psychiatric disorders, including addiction. The brain’s medial prefrontal cortex (mPFC) is important for social behavior, but the mechanisms by which mPFC neurons code real-time social exploration remain largely unknown.
An international team led by NIDA scientists used miniature fluorescence microscopes to record calcium activities from hundreds of excitatory neurons in the mPFC in the brains of mice as they interacted socially with other mice. The animals were given a series of social-behavior tests in the absence or presence of the psychedelic drug phencyclidine (PCP), which alters social behaviors. Findings revealed that ensembles, which are distinct and dynamic neural groups, are tuned on or off to social exploration, and dysfunctions in these ensembles are associated with abnormal social exploration elicited by PCP. These findings underscore the importance of mPFC “on” and “off” neural ensembles for proper social-exploratory behavior and pave the way for future studies related to neural-circuit dysfunctions in psychiatric disorders. (NIH authors: B. Liang, L. Zhang, G. Barbera, W. Fang, J. Zhang, Y. Li, and T. Lin, Neuron 2018; DOI:10.1016/j.neuron.2018.08.043)
[FROM NIDA SCIENCE HIGHLIGHTS, 10/9/18]
This page was last updated on Wednesday, April 6, 2022
