NIH Research Festival: Concurrent Symposia
BENCH-TO-BEDSIDE HOMERUNS

The NIH Clinical Center (CC) has been home to transformative stories of researchers who successfully took an idea from the bench to the bedside. Some of the NIH researchers who were beneficiaries of the CC’s Bench-to-Bedside and Back Awards were on hand to retell their homerun stories.
Peter Pinto’s team (National Cancer Institute, NCI) hit a homerun when they used high-resolution magnetic-resonance imaging transrectal ultrasound to diagnosis advanced prostate cancer and to distinguish aggressive high-grade prostate lesions from indolent tumors, which can be watched rather than requiring aggressive surgery.
National Human Genome Research Institute researchers Charles Venditti and Irini Manoli scored homeruns with their work on methylmalonic acidemia (MMA), an inherited disorder in which the body is unable to process certain proteins and fats properly. Venditti described how, in knockout and transgenic mice with MMA, the team could predict the severity of the disease—by using stable-isotope breath testing to screen for the enzyme methylmalonyl coenzyme A mutase (MUT)–and then use a variety of gene-delivery approaches to rescue the unstable metabolic symptoms. The team developed a similar breath test for humans with MMA as a “safe and noninvasive estimate of in vivo MUT enzymatic activity,” said Manoli. The test could provide an outcome parameter for gene, cell, or enzyme therapies for the disorder.
Yet another homerun was hit by Katherine Meilleur (National Institute of Nursing Research), who is studying common skeletal myopathies caused by mutations in the RYR1 gene. In a clinical trial, she discovered that patients suffering from RYR1-related myopathies showed increased “oxidative stress” as measured by concentrations of the antioxidant glutathione in plasma.
And the homeruns kept coming. Jack A. Yanovski (National Institute of Child Health and Human Development, NICHD), who pointed out that several genes “confer the potential for obesity,” hit a homerun with his discovery that mutations in the mc3r gene predisposed children to obesity. The mc3r gene codes for the protein melanocortin 3 receptor. In mouse studies, he found that the deletion of mc3r induced a decrease in lipid oxidation and an increase in adipose deposition in mesenchymal stem cells.
FARE Award winner Reema Railkar’s (NCI) homerun resulted from her use of quantitative high-throughput screening to identify novel therapies for bladder cancer. She described how the candidate drug flavopiridol was “highly effective” at inhibiting aggressive bladder cancer cells.
The “From Insight to Therapy: Bench-to-bedside Homeruns” session was co-chaired by John I. Gallin (Clinical Center) and Constantine A. Stratakis (NICHD).
THE MIGHTY MICROBIOME: DISTINGUISHING FRIENDS FROM ENEMIES
In the past, microbes were considered enemies to human health. Recent discoveries, however, have highlighted the key role that the human-associated microbiome plays in promoting health. Several intramural investigators discussed how their explorations of microbial communities have shed light on the etiology of disease and have helped them design interventions to promote health and treat disease.
Julie Segre (National Human Genome Research Institute, NHGRI) investigates how bacteria and other microbes that constitute the skin microbiome contribute to health and how changes in them can lead to chronic skin disorders such as eczema and psoriasis. Surprisingly, she found that the skin microbiome differs less among individuals than among microenvironments in the same person: dry, moist, and sebaceous.
The lung microbiome is the focus of Curtis Harris’s research (National Cancer Institute). He examined nearly 400 samples—frozen lung-cancer tumor samples and controls—and found an increase in Variovorax and Streptococcus bacteria in the tumors. He said that mechanistic studies must be performed to determine the role of the two taxa in cancer.
The oral cavity is second only to the gastrointestinal tract in the diversity of the microbial community. Niki Moutsopoulos (National Institute of Dental and Craniofacial Research) studies periodontitis, a microbial-stimulated disease that leads to the destruction of tooth-supporting structures. She examined the oral microbiome of patients with leukocyte-adhesion deficiency 1 (LAD1) and severe periodontitis, and she showed that the dysbiotic microbial communities in LAD1 may stimulate a local inflammatory response.
FARE Award winner Dennise A. de Jesus-Diaz (National Institute of Allergy and Infectious Diseases, NIAID), who is characterizing the microbiota of pediatric patients during a norovirus episode and recovery, observed an increase in proteobacteria in the intestinal microbiota of patients during norovirus infection. She is currently working on a new intestinal enteroid cell-culture system to incorporate secondary metabolites produced by bacteria.
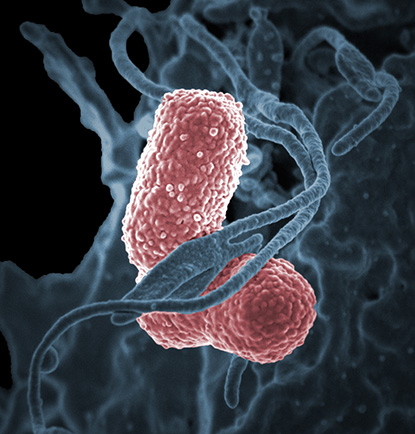
CREDIT: NIAID
NIH researchers are investigating multi-drug-resistant bacteria such as Klebsiella pneumoniae that can cause severe hospital infections. Shown: A human neutrophil interacting with Klebsiella pneumoniae (pink).
Multidrug-resistant microbes are also being investigated by intramural researchers. In 2011, a strain of carbapenem-resistant Klebsiella pneumonia colonized 18 patients at the NIH Clinical Center, with seven deaths attributed to the infections. Tara N. Palmore (NIAID) described how she collaborated with Segre’s team of researchers, who used advanced genomic technologies to hunt down and contain the bacteria. She even did a study to determine whether health-care workers had been colonized by the microbes and were possibly spreading them. Fortunately, she found no evidence of colonization.
With our growing knowledge of the human microbiome and advances in bioinformatics, we are now able to look at more-complex infection models and leverage this information to improve treatment and patient care.
The “Zen of Microbiota” session was chaired by Julie Segre (NHGRI).
TELL ME YOUR EXPOSURES AND I’LL TELL YOU WHO YOU ARE
Exposure to toxic environmental chemicals and agents while in utero and other developmentally sensitive times can have negative effects on later health outcomes in children, adults, and even future generations. Several intramural scientists who are exploring this field of developmental origins of health and disease (DOHaD) highlighted new evidence for DOHaD in human health outcomes as well as current data regarding the underlying mechanisms.
Using data from epidemiologic studies, Edwina Yeung (National Institute of Child Health and Human Development, NICHD) showed that obesity during pregnancy affects not only the mother’s health but also the early development of her child. Maternal obesity is associated with delays in the child’s motor development. Paternal obesity is related to delays in the child’s social functioning. Additionally, children with two obese parents can experience some lag in problem-solving skills.
In another epidemiological study, Martha Linet (National Cancer Institute) showed that repeated diagnostic X-ray exams during childhood and adolescence could result in a higher risk of breast cancer in adulthood. Moreover, children and teenagers who were exposed to radiation from atomic bombs or the catastrophic Chernobyl nuclear power plant accident in 1986 (in the Ukraine) are more susceptible to the development of solid tumors and thyroid cancers, respectively.
Epigeneticists are also conducting studies in an attempt to understand the molecular drivers of DOHaD. The researchers focus on how the epigenome, a myriad of chemical modifications to the DNA and DNA-associated proteins in a cell, is affected by environmental exposures or disease. Bruce Howard (NICHD) discussed how defects in the maintenance of epigenetic structures (or failures in programmed transitions, especially in the perinatal period) may underlie common developmental disorders and age-related diseases. He pointed out that certain genomic regions are more susceptible to altered epigenetic states that can lead to neurodegenerative diseases.
FARE Award winner Katryn Harwood (National Institute of Diabetes and Digestive and Kidney Diseases) is trying to understand the role of the O-linked beta-N-acetylglucosamine (O-GlcNAc) cycling enzymes in the epigenome. O-GlcNAc is an intracellular carbohydrate that modifies proteins in the nucleus and cytoplasm. Harwood proposes that the balance between the cycling enzymes O-GlcNAc transferase (which attaches O-GlcNAc to the protein) and O-GlcNAcase (which takes O-GlcNAc away from the protein) plays an important role in turning on and off transcription processes that control germ-line maintenance and embryo viability.
The timing of exposures can have different effects, according to Carmen Williams (National Institute of Environmental Health Sciences NIEHS), who talked about how estrogenic endocrine disruptors alter the ability of the female reproductive-tract environment to support fertilization and embryo development. A female exposed prenatally to estrogen is more likely to have a higher risk of malformations in her reproductive tract as an adult, Williams explained. On the other hand, mouse studies show that neonatal exposure can cause epigenetic changes that modify gene expression and increase the risk for uterine cancer.
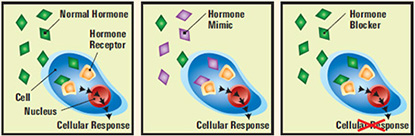
CREDIT: NIEHS
NIEHS scientist Carmen Williams talked about how estrogenic endocrine disruptors alter the ability of the female reproductive-tract environment to support fertilization and embryo development. Illustration shows that when absorbed in the body, an endocrine disruptor can decrease or increase normal hormone levels (left), mimic the body's natural hormones (middle), or alter the natural production of hormones (right).
The “Lasting Legacies: Long-term Effects of Early Developmental Exposures” session was chaired by Carmen Williams (NIEHS).
The Trigger of Chronic Disease: Inflammation
Inflammation can be protective when the body is reacting to harmful pathogens or irritants. But when inflammation becomes chronic or dysregulated, it can evolve into a pathophysiological response as is seen in chronic autoimmune, neurodegenerative, fibrotic, and allergic diseases and even in some cancers. Several NIH scientists have taken on the challenge of deciphering the mechanisms that trigger inflammation, and a few presented their work at a special symposium held during the Research Festival.
Skin: Many types of bacteria colonize healthy skin, but this diversity is lost in atopic dermatitis patients’ skin, which is dominated by Staphylococcus aureus. Using mouse models, Keisuke (Chris) Nagao (National Cancer Institute, NCI) discovered that S. aureus was the driving force behind the atopic inflammation and that the absence of the enzyme ADAM-17 impaired normal regulatory signaling. He is trying to understand why the overgrowth of S. aureus occurs and what the immunological pathways are downstream.
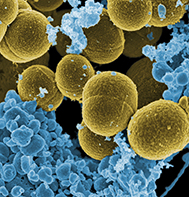
CREDIT: NIAID
Many types of bacteria colonize healthy skin, but this diversity is lost in atopic dermatitis patients’ skin, which is dominated by Staphylococcus aureus (shown). NCI researcher Chris Nagao discovered that S. aureus was the driving force behind atopic inflammation and is trying to understand why the overgrowth of the bacteria occurs.
Eyes: Inflammation plays a role in the development of age-related and other degenerative eye diseases. Kapil Bharti (National Eye Institute) is investigating how damage to the outer blood-retina barrier (made by the retinal pigment epithelium—RPE—and endothelial cells) causes macular edema (inflammation and fluid accumulation in the eye). He has created a novel model of macular edema by culturing RPE and endothelial cells that are differentiated from patient-specific induced pluripotent stem cells in microfluidic tissue chips. He is trying to discover the signaling pathways that cause the condition. Bharti’s team hopes their research will be useful for both preclinical and clinical studies related to inflammatory diseases that affect the human retina.
Liver cancer: Metabolic events control T-cell immunity and inflammation in liver cancer, according to Tim Greten (NCI). He compared risk factors for viral hepatitis and nonalcoholic liver disease–related inflammation and found commonalities in T-cell infiltration, immune signature, and natural-killer-cell function. He also found that a fatty diet induced fatty-liver buildup. He hopes that a better understanding of the underlying inflammatory mediators will advance research on inflammation-related tumor development.
Rebound inflammation: Few therapies exist for fibrotic disease such as asthma, liver cirrhosis, cardiovascular disease, idiopathic pulmonary fibrosis, Crohn disease, and ulcerative colitis. Thomas Wynn (National Institute of Allergy and Infectious Diseases, NIAID) discussed ways to improve antifibrotic therapy by fighting rebound inflammation (inflammation that develops in response to antifibrotic drugs). His group was the first to demonstrate a central and indispensable role for interleukin-13 (IL-13) in the development of fibrosis and has hypothesized that any intervention that disrupts critical steps in the IL-13 response might emerge as a viable therapeutic strategy for fibrosis.
Lung: Second-time FARE Award winner Seddon Thomas (National Institute of Environmental Health Sciences) is doing research to better correlate inflammatory asthmatic reactions with environmental causes. She summarized her research on the gene MYD88. This gene codes for the protein myeloid-differentiation primary response 88, which helps orchestrate the lung’s immune responses to inhaled allergens.
The “Rings of Fire: Inflammation at the Intersection of Chronic Diseases” session was chaired by Thomas Wynn (NIAID).
PRECISION MEDICINE NOW
NIH researchers are laying the groundwork for precision medicine—an arena that takes into account individual variability in genes, environment, and lifestyle. Advances in DNA-sequencing technology and in sequencing the human genome have made it possible for scientists to identify genetic mutations and then investigate their effect on disease risk and progression. At the symposium, several intramural researchers reported on their work using this precision-medicine approach to investigate a range of diseases.
Richard Siegel (National Institute of Arthritis and Musculoskeletal and Skin Diseases, NIAMS) explained how genetic variants that alter the expression of tumor necrosis factor (TNF)-pathway components can affect the risk of developing autoimmune and autoinflammatory diseases such as Behçet disease, Crohn disease, multiple sclerosis, primary biliary cirrhosis, rheumatoid arthritis, and ulcerative colitis.
Leslie Biesecker (National Human Genome Research Institute, NHGRI) outlined a framework for understanding genomic medicine in the context of the Precision Medicine Initiative, announced by President Obama in 2015. Biesecker directs ClinSeq, an NHGRI clinical study launched in 2007, that explores how genetics can inform patient health and medical diagnosis and improve genetic-data management and utilization. He described case studies and highlighted projects aimed at improving genomic-data analysis and data sharing.
Ivona Aksentijevich’s (NHGRI) use of genomic sequencing led to the discovery of two autoinflammatory diseases—haploinsufficiency of A20 and otulipenia—and the identification of a promising therapeutic approach that involves inhibiting TNF-cytokine signaling.
Javed Khan’s (National Cancer Institute, NCI) work also points to the clinical feasibility of using DNA sequencing to inform therapeutic approaches for treating relapsed and refractory tumors in children. He identified many genetic mutations in a clinical genomics study of pediatric cancer patients. In addition, he described NCI’s new “ClinOmics” program that enables genome-guided precision therapies for adults and children with cancer.
The session was concluded by FARE Award winner Ngoc-Han Ha (NCI), who presented her discovery of the role played by the circadian rhythm gene Arntl2 in determining metastatic susceptibility and progression in breast-cancer patients.
NIH expects personalized precision medicine to play a significant role in the tailoring of treatments for human disease in the future.
The “Precision Medicine Now: The Power of NIH Patient Cohorts to Wed Genotype with Phenotype” session was chaired by Richard M. Siegel (NIAMS).
CAN BIG DATA HELP US ANSWER BIG QUESTIONS?
With the decreasing costs and increasing speeds of sequencing technology, scientists are rapidly generating large volumes of data. Advances in computational biology may be able to help us interpret this overwhelming wealth of information. Several NIH investigators spoke at the Research Festival about using “big data” in their work.
José Faraldo-Gómez (National Heart, Lung, and Blood Institute) uses computer modeling to study membrane-associated proteins. In a simulated membrane environment, a computer can calculate the force on every single atom to determine the movements of each. A supercomputing infrastructure enables scientists to calculate in seconds what would have once taken a single processor decades to complete. Faraldo-Gómez has used supercomputing modeling to show that the enzyme that creates ATP drives the membrane perturbations that give the mitochondrion its distinct folded appearance. The folds increase the organelle’s surface area, enabling the cell to generate more ATP in a smaller volume. ATP is a complex macromolecule that powers almost every activity of the cell.
Jianxin Shi (National Cancer Institute, NCI) and colleagues performed an integrative genomic analysis for lung adenocarcinoma (LUAD) patients in the NCI EAGLE (Environment And Genetics in Lung Cancer Etiology) study. This study identified several driver genes in LUAD including two novel ones. Interestingly, though, 25 percent of the patients surveyed did not have mutations in existing driver genes, suggesting that novel mechanisms were behind many of these cancers. The study also showed that the number of somatic mutations and certain other mutation types was associated with increased risk of distant metastasis.
Can big data help us determine the optimal dose, timing, or drug combination for treating cancer patients? To answer this question, Orit Lavi (NCI) is exploring how redundancy causes problems in chemotherapy. Redundant elements are those that act in the same biologic or dynamic manner and can compensate for each other if necessary. For example, targeting an oncogene by inhibiting one of its transcriptional activators is useless if there are other activators that can take over. She argues that the integrated computational approaches of mathematical modeling and statistical analysis of information—on tumor heterogeneity, drug resistance, and redundant functions—may enable us to determine the optimal course of treatment.
FARE Award winner Jessica Petrick (NCI) studies the relationship between weight changes during a person’s lifetime and two types of cancer that have been increasing: esophageal and gastric cardia adenocarcinoma. Using data from two large cancer cohort studies, Petrick found that individuals who were overweight as young adults and obese later in life were at the highest risk for these cancers. Future work may look into whether weight gain between childhood and young adulthood has any effect on cancer risk.
The “Making Sense of Greek Letters and Too Many Numbers in the Age of Big Data” session was chaired by Stephen Chanock (NCI).
This page was last updated on Tuesday, April 12, 2022
