NIH Research Festival: Celebrating 30 Years
A Taste of NIH Intramural Research and More
A taste of NIH research. That’s what the Research Festival is all about. And for the past 30 years, the festival has provided a lively venue for intramural scientists to share their work with their peers.
“We had a lot of enthusiasm from scientists in organizing the sessions,” said festival co-chair Andrew Griffith, the scientific director of the National Institute on Deafness and Other Communication Disorders.
The festival, which was held on September 14–16 in the Clinical Center, featured three plenary sessions on some of NIH’s cutting-edge research—on superenhancers, clinical imaging, and cell-based immune therapies. The three days were also filled with concurrent symposia, poster sessions (including one for institute directors and scientific directors), special exhibits on intramural resources, the Green Labs Fair, the FARE Awards ceremony, demonstrations on three-dimensional printing and virtual reality, tours of the National Library of Medicine and the Clinical Center, the Technical Sales Association exhibit tent show, and more.
“A highlight was having [NIH Director Francis] Collins speak and present on his science,” said festival co-chair Ann Cashion, who’s the scientific director of the National Institute of Nursing Research. Collins spoke at the superenhancer plenary.
The poster sessions were another highlight and gave trainees a chance to shine. “Our trainees and fellows do exceptional work,” said Cashion. “It’s wonderful to see those who are going to replace me showing such promise and such unique new ways of thinking.”
Following are articles on the plenary sessions as well as the concurrent symposia. The plenary sessions were also videocast and are archived online.
Plenary Sessions
FIRST ON THE SCENE: INTRAMURAL RESEARCHERS TALK SUPERENHANCERS
Research Festival Plenary Session I
Although the Research Festival itself is celebrating its 30th year, the topic for the opening plenary session of 2016 has only just celebrated its 3rd birthday! With a potential role in helping conditions ranging from diabetes to cancer to infectious diseases, the newborn field of superenhancer research seems to have a great big future in front of it.

PHOTO BY ERNIE BRANSON
NIH Director Francis Collins kicked off the 2016 Research Festival with a talk on his research on stretch enhancers, noncoding regions of the genome that enhance gene expression from a distance.
In 2013, two different groups investigating epigenetics discovered that certain multi-kilobase noncoding regions of the genome seemed not only to be able to enhance gene expression from a distance but also to powerfully act on genes in specific cell types. One of these groups was led by Francis Collins, NIH director and head of the molecular genetics section in the National Human Genome Research Institute.
Collins opened the plenary session by telling the story of how his group came to discover what he terms “stretch enhancers,” a slightly broader category of strong enhancers that includes superenhancers. While looking for the mechanism for how genetic risk factors influence type 2 diabetes, his team found that the vast majority of risk-producing single-nucleotide polymorphisms (SNPs), discovered by genome-wide association studies, fell in noncoding regions of the genome. “If you really want to understand type 2 diabetes on a functional basis, and how genetic risks play out,” he said, “you have to roll up your sleeves and say, ‘We’re going to get into epigenetics.’”
And so they did. Using ChromHMM software to analyze epigenetic data, Collins’s group found that many of their diabetes-associated SNPs fell in certain long stretches of noncoding DNA with the histone marks characteristic of enhancers. Then they used a method called ATAC-seq to identify transcription-factor binding sites, within those regions, that are altered in the presence of a genetic risk variant. In so doing, they discovered multiple examples of motifs for the RFX (regulatory factor X) family of transcription factors. RFX “is much more of a master transcription factor than we had realized,” said Collins. In fact, this search led them to RFX6, which seems to be closely tied to the special functions of pancreatic islet beta cells (which store and release insulin) that are critical to the prevention of diabetes.
CREDIT: RAFAEL CASELLAS, NIAMS
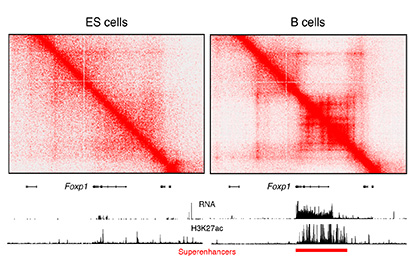
The image shows increased architectural interactions, as determined by Hi-C, associated with the Foxp1 superenhancer in B cells (right). In ES cells (left), the Foxp1 gene is silent (no RNA expression). Superenhancers are visualized as genomic domains enriched for H3K27Ac (a chromatin mark).
In the second talk, Rafael Casellas (National Institute of Arthritis and Musculoskeletal and Skin Diseases), joined by Erez Lieberman-Aiden from Baylor College of Medicine (Houston), laid out how a particular feature—specifically, the formation of long loops in the genome—is critical for superenhancers to do their work. When circulating B cells are activated by antigens, Casellas said, the process that leads to the production of antibodies involves a massive amplification of the transcriptome (the set of all messenger RNA molecules in a cell or population of cells). While examining the genome of these cells with super-resolution stochastic optical-reconstruction microscopy, Casellas noted that clusters of nucleosomes rapidly decompacted into monocucleosome fibers. This observation got him looking at how the architecture of chromatin changes when superenhancers go to work.
Using a technique called chromosome conformation capture using high-throughput sequencing (Hi-C), which detects chromatin interactions in the mammalian nucleus, Casellas and Lieberman-Aiden could spot long-range patterns in chromatin. These patterns “have become associated with many magic powers,” said Lieberman-Aiden.
In Hi-C maps, bright spots or stripes on the edges of squares indicate loops that form between flanking CTCF-binding sites, which act as insulators between domains of gene activiation.
“I love superenhancers because they’re the most striking examples of stripes we find in the genome,” said Casellas. “These beautiful constellations that we see in these [Hi-C] maps” can stretch for long distances. The longest of these superenhancer stripes is the one that includes the immunoglobulin heavy-chain locus—it has more than 100 CTCF-binding sites and stretches more than 2 million bases along the genome.
But how does the presence of these regions translate into changes in expression? In the final presentation of the session, Keiko Ozato (National Institute of Child Health and Human Development) discussed bromodomain-containing protein 4 (BRD4), a protein that binds acetylated histones such as those at superenhancer sites. BRD4 “is at the center of superenhancer research,” said Ozato. In macrophages, she explained, entire categories of expression programs that are activated after stimulation—such as inflammatory proteins, responses to injury, and other immune responses—are governed by superenhancers and most require BRD4, making proteins such as BRD4 master controllers for whole-gene expression programs.
“Superenhancers are diverse and fluid” and can potentially regulate a vast array of cellular programs, said Ozato. For a field still in its infancy, research on superenhancers seems to be quickly narrowing the gap between genome, gene expression, and cellular identity and function.
To view a videocast of the 2016 Research Festival’s Plenary Session I, “Super Enhancers in Cell Identity and Disease,” held on September 14, go to https://videocast.nih.gov/launch.asp?19852.
CLINICAL IMAGING: DETECTING BIOMARKERS FOR SPECIALIZED PATIENT CARE
Research Festival Plenary Session II
The NIH intramural program has helped to pioneer the field of clinical imaging, from the earliest development of magnetic-resonance imaging (MRI) technology and Louis Sokoloff’s now-classic positron emission tomography (PET) scan techniques to map and measure brain function, to recent breakthroughs such as MRI-guided closed-chest heart repair. This plenary session highlighted the latest research of three NIH scientists who are extending this tradition of clinical-imaging advances: Xiaoyuan (Shawn) Chen’s (National Institute of Biomedical Imaging and Biomedical Engineering), who discussed his work with nanotheranostics; Jessica Gill (National Institute of Nursing Research), who presented her discoveries of mechanisms underlying differential responses to traumatic brain injury; and PET expert Robert Innis (National Institute of Mental Health), who talked about his most recent method for imaging proteins as a biomarker for neuroinflammation.
CREDIT: SHAWN CHEN AND JIBIN SONG, NIBIB
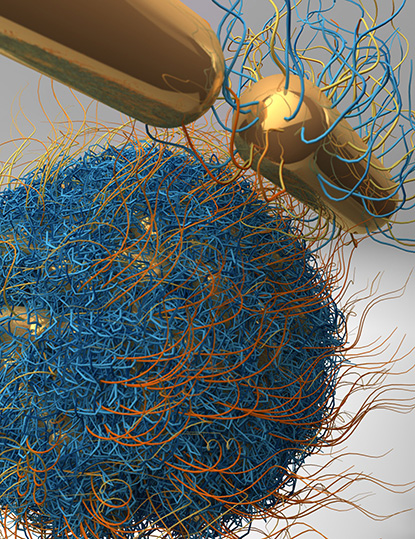
Shawn Chen’s lab is engineering nanoparticles to carry drugs to tumors. When targeted with a laser, the vesicles release their payload. Shown: A nanovesicle assembled from individual gold nanorods; the hairy-looking material is a polymer coating on the surface of the vesicle.
Chen’s lab is developing molecular-imaging tools that can help with the early diagnosis of disease, monitor response to therapy, and guide drug discovery and development. His nanotheranostics work—the integration of diagnostic and therapeutic function using nanotechnology—involves packaging drugs inside tiny, gold, star-shaped nanoparticles called AuNanostars and delivering them straight to where they’re needed in a patient. These AuNanostars can be engineered to accumulate within a tumor and then, when targeted with a laser, release their drugs. Within hours of delivering their payload, they collapse and are secreted from the body. Chen is hoping to collaborate with the FDA and pharmaceutical companies to get his lab’s inventions into mainstream medicine.
Neuronal imaging is one of the tools that Gill is using to improve methods for identifying and helping patients at high risk for psychological and neurological impairments after a concussion or other kind of traumatic brain injury. She is using the presence of tau proteins, commonly associated with Alzheimer disease (AD), as a biomarker for brain injury. The amount of the protein found in blood or sweat is positively correlated with the severity of the injury.
Although the tau proteins are present in such small concentrations that they are nearly undetectable, Gill’s group has tweaked the single-molecule arrays method to amplify the protein’s detection up to 300-fold. This method for detecting tau can be used outside the clinic, for example, on a football field or a battlefield, and enables clinicians to predict which patients can resume their activities safely after a head injury and which are at increased risk of developing chronic neurological deficits and will need additional interventions.
PET imaging to detect inflammation biomarkers may help physicians more accurately diagnose and treat patients with such neuroinflammatory disorders as depression and AD, according to Innis. Until now, diagnosing and treating neurological disorders was based on trial and error, but Innis’s imaging method would eliminate the guesswork. He has identified several biomarkers including a new line of radioligands that can be detected with PET. Innis is currently collaborating with pharmaceutical companies to bring this technology to the clinic.
To view a videocast of the “New Insights through Clinical Imaging” plenary session, held on September 15, go to https://videocast.nih.gov/launch.asp?19854.
HARNESSING THE IMMUNE SYSTEM TO FIGHT DISEASE
Research Festival Plenary Session III
Immunotherapy is a centuries-old concept of stimulating the immune system to fight noninfectious disease. It has become a viable treatment thanks to the pioneering work of National Cancer Institute (NCI) researchers, that of Michael Potter on monoclonal antibodies in the 1960s and Robert Gallo on interleukin-2 (IL-2) in the 1970s. This plenary session featured three NIH scientists who built on these advances and have been able to harness the immune system to fight cancer and other chronic, noninfectious diseases: Steven Rosenberg (NCI), who was the first to recognize the potential of IL-2 and apply it as a novel anti-cancer agent in 1984; Nicholas Restifo (NCI), who discussed new immunotherapies for patients with advanced cancer; and John Tisdale (National Heart, Lung, and Blood Institute), who presented his research on combining hematopoietic stem-cell methods with immunotherapy for the treatment of the genetic-based sickle-cell disease.
CREDIT STEVEN ROSENBERG, NCI
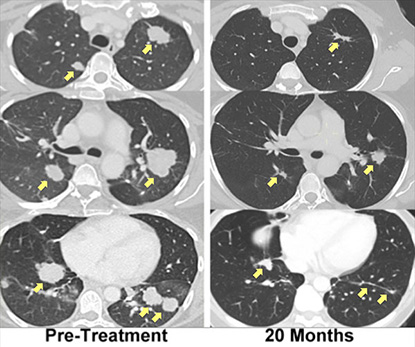
A patient with metastatic bile duct cancer, treated with immunotherapy using her own mutation-specific T cells, has experienced regression of her metastatic lung and liver tumors that had been ongoing for more than two years.
Rosenberg, widely regarded as the father of immunotherapy, developed the first effective immunotherapies in patients with advanced cancer and was the first to successfully import genes into humans. In his talk, “Cells as Drugs: A Personalized Immunotherapy for Cancer,” he emphasized the ability of human lymphocytes to recognize unique cancer antigens and how antitumor T-cell receptors can be exploited to develop new cell-transfer immunotherapy treatments for cancer.
He explained the process of adoptive cell therapy, which uses tumor-infiltrating lymphocytes, and how lymphodepletion before infusion dramatically improves the effect of transferred cells by eliminating cells that endogenously consume cytokines for T-cell growth. The identification and targeting of mutations unique to each cancer has the potential to extend cell therapy to patients with epithelial cancers. Autologous lymphocytes genetically engineered to express T-cell receptor or chimeric antigen receptors can mediate the regression of metastatic cancers. Rosenberg’s studies on cell-transfer immunotherapies has resulted in durable, complete remission in patients with metastatic melanoma. Recently, he developed chimeric antigen-receptor therapy and T-cell–receptor gene-engineered cell therapy to fine-tune and turbocharge the killing of cancer cells.
Restifo, one of Rosenberg’s former postdocs and a pioneer in the use of T-cell–based immunotherapies, was the first to identify myeloid-derived cell substances that impair antitumor T-cell responses in humans. In his talk “Novel Elements of Immune Suppression within the Tumor Microenvironment,” he discussed the immunology of oxygen and potassium and how they can be used to destroy cancer. His lab is currently working on reprogramming T cells to induce curative responses in patients with metastatic cancer.
Cell-based immune therapy is also being explored as a way to treat sickle-cell disease. In his talk “Exploiting Hematopoetic Stem Cells for Treating Genetic Disease: Sickle-cell Disease,” Tisdale described his work combining hematopoietic stem-cell transplant methods with immunotherapy. Usually, a patient’s immune system must be suppressed so transplanted cells and tissue aren’t rejected, and preventing rejection can mean a lifetime of taking immunosuppressant drugs. But Tisdale may have a solution: He found that, in adult patients, hematopoietic stem cells from matched sibling donors could be successfully transplanted without having to destroy the patient’s immune system.
To watch a videocast of the 2016 Research Festival’s “Cell-based Immune Therapy” plenary session, held on September 16, go to https://videocast.nih.gov/launch.asp?19862.
More Photos

CREDIT: ERNIE BRANSON
Postdoc Brianna Bingham (National Institute of Diabetes and Digestive and Kidney Diseases) discusses her research on insulin resistance in African American women.

CREDIT: ERNIE BRANSON
Not only did the scientific directors and institute directors participate in the poster session, but some of them showed off their baking talents. Festival co-chair Andy Griffith’s apple pastries won the bake-off. Pictured from left: Stephen Channock (one of NCI’s scientific directors), Jackie Roberts (Festival coordinator), and Richard Wyatt (Deputy Director, Office of Intramural Research).

CREDIT: ERNIE BRANSON
Even the scientific directors and institute directors displayed posters at the 2016 Research Festival. Tom Mistelli (one of the scientific directors at NCI) won for his poster on "Repression of the antioxidant NRF2 pathway in premature aging.” Pictured: Michael Gottesman (NIH Deputy Director for Intramural Research) points out details on his poster on P-glycoprotein (P-gp), a multi-drug transporter that was identified and cloned in his lab in 1986. P-gp has been shown to contribute to multidrug resistance in cancer and to be a key element in pharmacokinetics and organ distribution of many drugs in current use.

CREDIT: ERNIE BRANSON
The contributions of research animals were recognized at a special ceremony on the Clinical Center’s south lawn during the Research Festival. On September 16, the NIH intramural animal research advisory committee and the IC animal program directors dedicated a plaque that commemorates “research animals and the NIH animal care-and-use community that have contributed to our exceptional biomedical research advances.” Pictured: NIH Director Francis Collins, Deputy Director for Intramural Research Michael Gottesman, retired animal program director Shelley Hoogstraten-Miller, National Eye Institute senior investigator Rachel Caspi, and director of the NIH Office of Animal Care and Use Terri Clark.
This page was last updated on Tuesday, April 12, 2022
