Research Briefs
NIAID: SOFT PALATE MAY BE KEY BREEDING GROUND FOR INFECTIOUS FLU
A collaboration between NIAID and MIT scientists has revealed that the soft palate, the soft tissue at the back of the roof of the mammalian mouth, may be important for flu transmission in humans. Historically, the soft palate has not been examined in animal models of influenza.
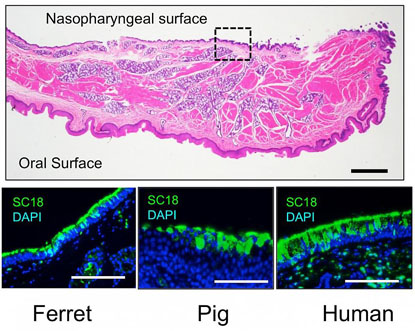
CREDIT: NIAID
Flu viruses enter cells by binding to sialic acids on surface glycoproteins. In ferrets, pigs, and people, the nasopharyngeal surface of the soft palate contains regions of densely packed long-chain alpha 2,6 sialic acid molecules (shown in green) where influenza viruses with airborne transmissibility can outcompete less transmissible viruses.
Flu infection starts when hemagglutinin (a substance that causes red blood cells to clump) proteins on an influenza virus bind to sialic acid (SA) molecules on the tops of chainlike proteins that thickly line tissue throughout the respiratory tract. Forms of the flu that infect birds bind to a type of SA called alpha 2,3 SA, whereas those that infect humans and other mammals bind to a type of SA called alpha 2,6 SA.
The scientists made four mutations in the hemagglutinin of the 2009 influenza pandemic strain, which was good at spreading from person to person. The intent of introducing the mutations was to make the virus preferentially bind to bird-type SA and, presumably, be less transmissible via air than the original virus. The engineered virus was then used to infect a group of ferrets, which are widely used as a model of human influenza infection. The researchers were surprised to find that the engineered flu virus was transmitted by the airborne route to uninfected ferrets just as well as the original nonmutated virus.
The researchers concluded that certain characteristics of the soft palate might provide a competitive advantage to more-contagious varieties of the flu, allowing them to reproduce rapidly in the area and outcompete other strains. This information could aid efforts to define the properties governing flu-virus transmissibility and predict which viruses are most likely to spark pandemics. (NIH authors: S.S. Lakdawaka, E.W. Lamirande, A.R. Shih, C.T. Hanson, L. Vogel, M. Paskel, M. Minai, I. Moore, M. Orandle, and K. Subbarao, Nature 526:122–125, 2015)
OD: NEW METHOD FOR MEASURING THE IMPACT OF SCIENTIFIC PUBLICATIONS
A working group in the NIH Office of the Director has created a new method for assessing the impact of scientific journal articles. The quality of a paper is commonly judged by examining the journal impact factor (JIF) of the journal in which it was published. The JIF reflects the average number of citations to all papers published in the previous two years in a journal, a measure of the journal’s importance that has many weaknesses and obviously, when used as a proxy for a specific article’s impact, underestimates the influence of a journal’s most important papers.
The working group’s alternative measure, called the relative citation ratio (RCR), compares the number of times a paper has been referenced with the number of citations received by a set of related papers. The team conducted an analysis of thousands of journal articles and determined that their RCRs were strongly associated with experts’ assessments of their quality. Moreover, the group found that only 11 percent of papers with a high RCR appear in journals with a high JIF, suggesting that the JIF misjudges the impact of 89 percent of influential papers.
Although the RCR metric will require further testing and fine-tuning before it is widely adopted, it nonetheless has the potential to help scientists, funding agencies, and policymakers more meaningfully evaluate scientific research. (NIH authors: B.I. Hutchins, X. Yuan, J.M. Anderson, and G.M. Santangelo, bioRxiv DOI10.1101/029629)
NCI, NIAMS, NHLBI, CC: NEW WAY FOR KILLER T CELLS TO COMMUNICATE
NIH researchers described for the first time an entirely new way in which anticancer killer T white-blood cells can directly communicate with one another to synchronize and coordinate their behavior. This behavior, which is reminiscent of quorum behavior observed in bees, ants, and microorganisms, allows T cells to function not as single actors but as an organized army moving in unison. A commentary that appeared in the same issue of the journal in which the NIH study was published, reported: “The clinical application of T cell immunotherapy depends on ex vivo modification and expansion of T cells for adoptive transfer. In preclinical models, the use of a purified, naive T cell subset enhances persistence and antitumor immunity; however, the majority of clinical studies rely on modification of mixed populations of T cells that contain only a small subset of highly functional T cells with less-differentiated phenotype. [The NIH researchers] uncover[ed] a Fas-mediated interaction between naive T cells and antigen-experienced T cells that drives differentiation and impairs adoptive immunotherapy. Further, [the study] show[ed] that blockade of Fas signaling enhances antitumor immunity and increases survival in a mouse model of melanoma. [The] work supports a growing body of evidence that the use of naive T cells enhances the efficacy of adoptive T cell therapy and suggests a new therapeutic strategy for preserving less-differentiated T cell populations.” The research is directly influencing current T-cell-therapy clinical trials (NCT02062359) being conducted at the Clinical Center. (NIH authors: C.A. Klebanoff, C.D. Scott, A.J. Leonardi, T.N. Yamamotoa, A.C. Cruz, R.M. Siegel, N.P. Restifo, and many others, J Clin Invest DOI:10.1172/JC181217; article: http://www.jci.org/articles/view/81217; commentary: https://www.jci.org/articles/view/85631)
NIDDK: PROTEIN SHUFFLES MICROORGANISM DNA
Many bacteria and microorganisms have an adaptive immune system known as the “clustered regularly interspaced short palindromic repeat” system, or CRISPR. This system allows for the recognition and destruction of invading viral DNA. NIDDK researchers have discovered that a protein related to one that enables CRISPR can relocate fragments of DNA.
The investigators studied Aciduliprofundum boonei, a microorganism that lives on the ocean floor. Before this research, it had been proposed that one of the organism’s CRISPR-associated protein 1 (Cas1) proteins—not part of the CRISPR—might be part of a new type of mobile DNA element. The investigators confirmed that this protein is in fact able to splice a specific piece of the organism’s DNA into new and random locations: a variation on a CRISPR ability.
The findings add to the understanding of how defense systems evolve and biochemical pathways develop and could have implications for genetic engineering of multicellular organisms. The investigators will next study whether this Cas1 protein can remove DNA from the original spot in the genome. (NIH authors: A.B. Hickman and F. Dyda, Nucl Acid Res 43:10576–10587, 2015)
NICHD, NCI: IMPROVING GENE THERAPY BY STUDYING HIV
NICHD and NCI researchers and outside colleagues have clarified how the human immunodeficiency virus (HIV) targets specific genes as the virus integrates into the human genome, the process that allows the virus to replicate. The team found that HIV integrates most often into genes that have a high density of introns—intervening sequences that are removed, or spliced, as genetic information is processed into readable instructions. After identifying nearly one million integration sites to find the virus’s preferred genes, the team discovered that many of these genes also have known roles in cancer progression.
The findings have important implications for gene therapy. Gene delivery relies on weakened viruses as vectors to convey the new genes into cells. In the search for a safe and efficient vector, researchers have tested many viruses. HIV, a lentivirus, is considered a better alternative to previous vectors, such as gamma retroviruses, which caused cancer in some patients.
Because lentiviruses are commonly used in current gene-therapy trials, the authors caution that gene recipients should be monitored carefully for overgrowth of treated cells that may eventually lead to cancer. The researchers hope that their findings will one day result in improvements in vector design to sidestep cancer risk without sacrificing the efficiency of gene delivery.
The team also clarified the function of lens epithelium–derived growth factor (LEDGF), a host protein that HIV hijacks in order to integrate its genetic material into human genes. The researchers found when LEDGF is removed from cells, HIV no longer preferred highly spliced genes. Also, in the absence of LEDGF, the splicing patterns of genes were altered. This result, along with other data, reveals that LEDGF is not just a facilitator for HIV; it also plays a role in gene splicing. (NICHD authors: Parmit K. Singh, James R. Iben, and Henry L. Levin; NCI authors: Andrea L. Ferris and Stephen H. Hughes, Genes Dev 29:2287–2297, 2015)
NIEHS: LIN28 IS A KEY PROTEIN AT WORK IN BREAST CANCER
The RNA-binding protein protein lin-28 (LIN28) plays a critical role in development timing and in cancer, but the molecular mechanisms underlying its role in promoting cancer has been poorly understood until now. A study by NIEHS scientists found that LIN28 uses diverse gene-regulatory mechanisms and functions as a master regulator of gene networks that modulate specific hallmarks of cancer. The findings, highlighted by journal editors in a “Spotlight” feature in Molecular Cell Biology, represent an important advance in basic science because it enhances our understanding of the mechanisms that lead to breast cancer.
The LIN28 protein occurs in two forms in mammals—LIN 28 homolog A (LIN28A) and LIN28B. LIN28A, through its regulation of splicing and gene expression, appears to produce a diverse family of proteins. The researchers propose that alternative splicing events produced downstream by LIN28 can affect specific breast-cancer subtypes and prognoses, a concept supported by their analysis of breast-cancer cells using RNA-Seq, RIP-Seq, and a custom computational method. Analyzing data from The Cancer Genome Atlas, the team found that LIN28 expression in the human epidermal growth factor receptor 2 (HER2) breast-cancer subtype was significantly higher than in other breast-cancer subtypes, although the reason for that overexpression remains unknown.
As a master regulator, LIN28 is a potential therapeutic target. The researchers pointed out that the suite of mRNAs whose expression is affected by LIN28 includes genes that function in cell metabolism, the immune response, cell proliferation, and cell-to-cell communication—processes that are classic hallmarks of breast-cancer biology. (NIEHS authors: J. Yang, B.D. Bennett, S. Luo, K. Inoue, S.A. Grimm, P.R. Bushel, H.K. Kinyamu, and T.K. Archer, Mol Cell Biol 35:3225–3243, 2015)
NIEHS: RGS2 PROTEIN HELPS PREPARE HEALTHY EGG-SPERM UNION
NIEHS researchers, collaborating with colleagues at three universities, have discovered that the RGS2 protein functions as a brake to suppress premature calcium release in eggs that are poised on the brink of development. While other researchers have shown that RGS2 plays an important role in regulating heart function and blood pressure, this is the first demonstration of the protein’s significant role in fertilization.
During the maturation process, the egg stores calcium, preparing it for fertilization. At fertilization, the sperm causes calcium to be released in the egg, turning it into a developing embryo. The mouse study showed that during maturation, the egg synthesizes RGS2, which suppresses calcium signaling. This safety mechanism ensures that the egg does not begin releasing calcium and start developing before the sperm arrives, as that would prevent the egg from merging with the sperm.
RGS2 is being studied as a therapeutic target for hypertension and other heart ailments. Understanding the role RGS2 plays in reproduction is important when considering the possible benefits and side effects of any new treatments, as well as understanding the impact that toxins might have on human fertility. (NIH authors: M.L. Bernhardt, E. Padilla-Banks, C.E. McDonough, and C.J. Williams. Development; doi: 10.1242/dev.121707)
NICHD: HUMAN TASTE MAY BE MORE DISCERNING THAN PREVIOUSLY THOUGHT

PHOTO COURTESY OF GRAEME LOWE AND THE JOURNAL OF NEUROSCIENCE
A new NICHD study contradicts the commonly held model of how the brain processes taste. The theory contends that all chemicals can be grouped into five different taste categories—bitter, sweet, salty, sour, and umami—that are each processed by distinct neural pathways. The researchers delivered small amounts of various molecules to the taste receptor neurons of moths and directly measured the cells’ responses. They found that the timing of when and the pattern in which the cells fired when exposed to a compound was unique for each chemical rather than identical for chemicals in the same taste category. Because the insect taste system is thought to be similar in many ways to the mammalian one, the results suggest that both insect and human brains may be capable of distinguishing the tastes of thousands of different substances. (NIH authors: S. Reiter, C.C. Rodriguez, K. Sun, and M. Stopfer; J Neurosci 35:12309–12321, 2015)
NIDCR: ACTIVATING BRAIN FIELDS FOR TASTE
An NIDCR intramural neuroscientist and colleagues at Columbia College of Physicians and Surgeons (New York) and Howard Hughes Medical Institute (Ashburn, Va.) have demonstrated that stimulating defined regions of the brain’s cortex can drive taste behavioral responses in the absence of bitter or sweet substances in the oral cavity. By using optogenetics to activate specific regions of the taste cortex, the neuroscientists fooled mice into thinking that bitter or sweet substances were tickling their taste buds. Remarkably, stimulating the bitter brain field provoked gagging and attempts to clean the mouth of a nonexistent bitter substance. In contrast, stimulating the sweet brain field elicited compulsive licking. The study also showed that activating the bitter brain field could mask attraction to a sweet substance, and stimulating the sweet brain field could mask aversion to a bitter substance.
The taste cortical fields not only are able to elicit sensation of sweet and bitter, but also are necessary for taste perception. When the scientists silenced the bitter fields with a glutamate receptor antagonist, animals could no longer reliably identify bitter substances. However, their ability to recognize sweet substances was not impaired. The research team also showed that the sweet brain fields are required for an animal to perceive sweetness but not bitterness. (NIH author: Nicholas J.P. Ryba, Nature 527:512–515, 2015)
NCI, NIDDK: NEW COMPOUND REDUCES WEIGHT OF OBESE MICE
NCI and NIDDK researchers and colleagues created a new compound, called glycine-beta-muricholic acid (Gly-MCA), that reduces the weight of obese mice by acting on their intestinal cells. Gly-MCA is a bile-acid-based drug that selectively limits the action of bile-acid receptor farnesoid X receptor (FXR), an important regulator of energy metabolism. Gly-MCA improves beige-fat biogenesis via the inhibition of FXR in intestinal cells. The researchers treated five mice with Gly-MCA in several experiments and showed that the drug reduced weight gain in obese mice and improved their metabolic functions. The new compound may contribute to the development of new therapeutic approaches for obesity-related metabolic disorders. (NIH authors: F. Gonzalez, C. Jiang, C. Xie, K.W. Krausz, J. Shi, C.N. Brocker, and O. Gavrilova, Nature Commun DOI:10.1038/ncomms10166)
NIDDK: BOOSTING ACTIVITY IN BRAIN REGION MAY AID WEIGHT LOSS
NIDDK researchers have demonstrated that applying transcranial direct current stimulation (tDCS) to the brain’s dorsolateral prefrontal cortex (DLPFC) can reduce appetite and promote weight loss. Previous work by the group had found less neuronal activity in the left DLPFC of obese individuals, and the current work confirms the involvement of this region in eating behavior.

CREDIT: MARCI GLUCK, NIDDK
NIDDK scientists found that stimulating a part of the brain can reduce appetite and promote weight loss. Pictured: The study’s computerized vending machine tracked the quantity and type of foods participants consumed.
Nine obese subjects completed two separate eight-day stays in NIDDK’s metabolic ward, during which they ate a weight-maintaining diet for the first five days. Before breakfast on the final three days of the first visit, five participants received cathodal tDCS to the left DLPFC, whereas during this period of the second visit, they received anodal tDCS to this area. Cathodal stimulation tends to suppress neuronal activity, and anodal simulation boosts it. The four control subjects received sham stimulation on both visits. After stimulation, all subjects were allowed to eat freely for the rest of the day from a vending machine that tracked the quantity and type of foods they consumed.
Subjects ate fewer calories and lost more weight during the three days they were given anodal stimulation than on the days they received cathodal stimulation. Specifically, they consumed less soda and dietary fats. The group given sham stimulation showed no such differences between visits, suggesting that increasing left DLPFC activity with tDCS may have clinical utility for the treatment of obesity. The results will next need to be replicated in a larger group of subjects. (NIH authors: M.E. Gluck, P. Piaggi, C.M. Weise, R.J. Schwartzenberg, M. Reinhardt, E.M. Wasserman, C.A. Venti, S.B. Votruba, and J. Krakoff; Obesity 23:2149–2156, 2015)
NIDDK, NHLBI: NATURAL COMPOUND PREVENTS OBESITY IN MICE
A team of NIDDK and NHLBI researchers has found that treatment with celastrol enabled normal-weight male mice to avoid obesity and metabolic dysfunction despite being fed a high-fat diet. Celastrol is a natural compound extracted from the roots of Tripterygium wilfordii (thunder god vine) and Celastrus regelii. The findings suggest that celastrol kept the mice lean by transforming their white fat, which stores energy, into brown-like fat, which burns calories, and by increasing mitochondrial capacity and muscle endurance.

Thunder God vine
Excess white fat is associated with type 2 diabetes and other serious metabolic consequences of overweight and obesity. On the other hand, brown fat consumes and dissipates calories to generate heat. The results suggest that by “browning” white fat, celastrol is critical in preventing obesity in mice. Further study is needed to determine the safety and effectiveness of the compound in people.
Previous research found that celastrol decreased weight in obese mice by enhancing the action tunof the appetite-controlling hormone leptin. The current study showed that lower doses of celastrol can prevent obesity without affecting hunger, suggesting different capabilities of the herbal extract that depend on initial weight of the mice. (NIH authors: M.A. Xinran, X. Lingyan, A.T. Alberobello, O. Gavrilova, A. Bagattin, M. Skarulis, J. Liu, T. Finkel, and E. Mueller, Cell Metab 22:695–708, 2015)
This page was last updated on Monday, April 25, 2022
