Research Briefs
CC: ADVANCES MADE IN DIAGNOSING CORONARY HEART DISEASE
NIH Clinical Center researchers recently published a study that could help health-care professionals better identify, diagnose, and treat coronary-artery disease more quickly and effectively. The disease accounts for 17 million deaths a year.
Over time, coronary arteries, which supply blood to the heart muscle, can develop a buildup of plaque that hardens and eventually calcifies, narrows the arteries, and can rupture, reducing the flow of oxygen-rich blood to the heart.
An indicator of the calcification of the heart's arteries can be seen through a coronary-artery calcium score from a computed tomography (CT) scan. However, plaque that is still noncalcified, or soft, is not included in that score. The researchers believed that this type of plaque could also indicate an increased risk for the disease. And they were right.
When researchers compared the calcium score of the hardened plaque with that of the noncalcified soft plaque, they found significant differences. Noncalcified plaque that had not yet hardened was associated with low-density lipoprotein (“bad cholesterol”), diabetes, and systolic blood pressure. These variables were not associated with the calcium score in low-risk, asymptomatic individuals. Age was also a strong predictor of the calcium score, but not of noncalcified soft-plaque buildup. These results suggest that analyzing the different types of plaque, both hard and soft, gives more information about the risk of coronary-artery disease than just the calcium score.
This new method of relying upon plaque analysis and not just coronary-artery calcium score of hardened plaque shows promise as a tool for measuring the total plaque burden. (CC authors: K. Rodriguez, A.C. Kwan, D. Vigneault, V. Sandfort, P. Pattanayak, M.A. Alhlman, M. Mallek, C.T. Sibley, and D.A. Bluemke, Radiology DOI:http://dx.doi.org/10.1148/radiol.2015142551)
(From http://clinicalcenter.nih.gov/about/news/newsletter.html)
NCI, NHLBI: HIGH-RESOLUTION 3D IMAGES REVEAL THE MUSCLE MITOCHONDRIAL POWER GRID
A new imaging study in mice overturned longstanding scientific ideas of how energy is distributed within muscles for powering movement. NCI and NHLBI scientists reported the first clear evidence that muscle cells distribute energy primarily by the rapid conduction of electrical charges through a vast, interconnected network of mitochondria—the cell’s “powerhouse”—in a way that resembles the wire grid that distributes power throughout a city. The study offers an unprecedented, detailed look at the distribution system that rapidly provides energy throughout the cell where it is needed for muscle contraction.
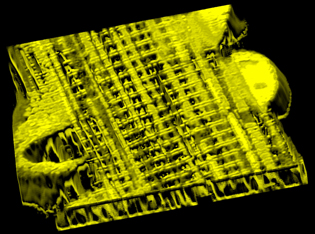

IMAGE BY BRIAN GLANCY AND CHRIS COMBS, NHLBI, NIH
This high-resolution 3D microscopic image shows a network of interconnected mitochondria within a mouse muscle cell.
The scientists collaborated in a detailed study of the mitochondria structure, biochemical composition, and function in mouse skeletal muscle cells. After using high-resolution three-dimensional (3D) images to reveal the structure of the mitochondrial power grid, the scientists then used specially designed optical probes to demonstrate that these mitochondrial “wires” were electrically conductive.
Using these probes, the researchers were able to demonstrate that most of the mitochondria were in direct electrical communication through the interconnecting network, demonstrating that the mitochondria are electrically coupled and can rapidly distribute the mitochondrial membrane voltage—the primary energy for ATP production—throughout the cell.
The study provides unprecedented images of how these mitochondria are arranged in muscle.
The use of focused ion-beam scanning electron microscopy, called FIB-SEM, played a critical role in unraveling the 3D architecture of mitochondrial networks in these muscle cells. The NCI scientists were the first to develop and use this imaging approach to image cells and tissue almost a decade ago.
Identifying this basic property of the muscle cell—how it distributes energy—has potential implications for disease diagnosis and treatment. In the future, scientists may use muscle biopsies or sophisticated noninvasive imaging techniques to determine how defects in mitochondrial networks affect different diseases. (NHLBI authors: Brian Glancy, Daniela Malide, Zu-Xi Yu, Christian A. Combs, Patricia S. Connelly, and Robert S. Balaban; NCI authors: Lisa M. Hartnell and Sriram Subramaniam, Nature 523:617–620, 2015. Link to article: http://www.nature.com/nature/journal/v523/n7562/full/nature14614.html#affil-auth)
NEI: IN BLINDING EYE DISEASE, TRASH-COLLECTING CELLS GO AWRY, ACCELERATE DAMAGE
Spiderlike cells inside the brain, spinal cord, and eye hunt for invaders, capturing and then devouring them. These cells, called microglia, often play a beneficial role by helping to clear trash and protect the central nervous system against infection. But a new study by NEI researchers shows that they also accelerate damage wrought by blinding eye disorders, such as retinitis pigmentosa. The findings suggest that microglia may provide a target for new therapeutic strategies aimed at halting blinding eye diseases of the retina.
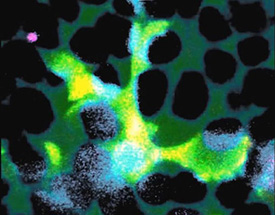
IMAGE BY WAI WONG, NEI
A microglial cell (green) extends spider-like arms to capture and consume rod photoreceptor cells (blue).
Retinitis pigmentosa, an inherited disorder that affects roughly 1 in 4,000 people, damages the retina, the light-sensitive tissue at the back of the eye. Research has shown links between retinitis pigmentosa and several mutations in genes for photoreceptors, the cells in the retina that convert light into electrical signals that are sent to the brain via the optic nerve. In the early stages of the disease, rod photoreceptors, which enable us to see in low light, are lost, causing night blindness. As the disease progresses, cone photoreceptors, which are needed for sharp vision and seeing colors, can also die off, eventually leading to complete blindness.
The investigators studied mice with a mutation in a gene that can also cause retinitis pigmentosa in people. In these mice, very early in the disease process the microglia infiltrate a layer of the retina near the photoreceptors, called the outer nuclear layer, where they don’t usually venture. The microglia then create a cuplike structure over a single photoreceptor, surrounding it to ingest it in a process called phagocytosis. The researchers caught this dynamic process on video. The whole feast, including digestion, takes about an hour.
Phagocytosis is a normal process in healthy tissues and is a key way of clearing away dead cells and cellular debris. However, in retinitis pigmentosa the researchers found that the microglia target damaged, living photoreceptors in addition to dead ones.
The researchers found evidence that photoreceptors carrying mutations undergo physiological stress, which then triggers them to secrete chemicals dubbed “find me” signals, which is like ringing a dinner bell that attracts microglia into the retinal layer. Once there, the microglia probe the photoreceptors repeatedly, exposing themselves to “eat me” signals, which then trigger phagocytosis. In response to all the feasting, the microglia become activated. That is, they send out their own signals to call other microglia to the scene, and they release substances that promote inflammation.
A clinical trial (NCT02140164) is already underway and recruiting participants at NEI to see whether the anti-inflammatory drug minocycline can block the activation of microglia and help slow the progression of retinitis pigmentosa. To watch a video of microglia eating rod photoreceptors, go to http://youtu.be/xXmUGYCi7rE. (NEI authors: L. Zhao, M.K Zabel, X. Wang, W. Ma, P. Shah, R.N. Fariss, H. Qian, and W.T Wong, EMBO Mol Med 7:989–1086, 2015)
NICHD: RESEARCHERS DESIGN PLACENTA-ON-A-CHIP TO BETTER UNDERSTAND PREGNANCY
NIH researchers and their colleagues have developed a “placenta-on-a-chip” to study the inner workings of the human placenta and its role in pregnancy. The device was designed to imitate, on a microlevel, the structure and function of the placenta and model the transfer of nutrients from mother to fetus. This prototype is one of the latest in a series of organ-on-a-chip technologies developed to accelerate biomedical advances.
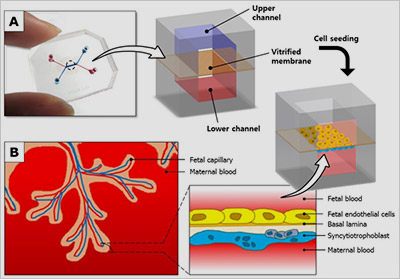
Figure: A placenta-on-a-chip microdevice: A) The device’s upper (blue) and lower (red) chambers are separated by a semi-permeable membrane. B) Researchers placed maternal cells in one chamber and fetal cells in the other. They then added glucose to the maternal-cell chamber and observed how it traveled through the membrane to the fetal-cell chamber.
The study was conducted by an interdisciplinary team of researchers from NICHD, the University of Pennsylvania, Wayne State University/Detroit Medical Center, Seoul National University, and Asan Medical Center in South Korea.
Studying the placenta in humans is challenging: it is time-consuming, subject to a great deal of variability, and potentially risky for the fetus. For these reasons, previous studies on placental transport have relied largely on animal models and on laboratory-grown human cells. These methods have yielded helpful information, but are limited in how well they can mimic physiological processes in humans.
The researchers created the placenta-on-a-chip technology to address these challenges, using human cells in a structure that more closely resembles the placenta’s maternal-fetal barrier. The device consists of a semipermeable membrane between two tiny chambers, one filled with maternal cells derived from a delivered placenta and the other filled with fetal cells derived from an umbilical cord.
After designing the structure of the model, the researchers tested its function by evaluating the transfer of glucose from the maternal compartment to the fetal compartment. The successful transfer of glucose in the device mirrored what occurs in the body.
The researchers hope that with further improvements, the technology may lead to better understanding of normal placental processes and placental disorders. (NICHD author: Roberto Romero, J Matern Fetal Neonatal Med DOI:10.3109/14767058.2015.1038518. Link: http://informahealthcare.com/doi/full/10.3109/14767058.2015.1038518)
NIDDK, USDA: NIH BODY-WEIGHT PLANNER ADDED TO USDA SUPERTRACKER FOOD AND ACTIVITY TOOL
The U.S. Department of Agriculture (USDA) and NIH have partnered to add the NIH Body Weight Planner (http://www.niddk.nih.gov/health-information/health-topics/weight-control/body-weight-planner/Pages/bwp.aspx) [SSR1] to USDA’s SuperTracker online tool (https://www.supertracker.usda.gov) [SSR2] as a goal-setting resource to help people achieve and stay at a healthy weight. Created in 2011, the SuperTracker tool empowers people to build a healthier diet, manage weight, and reduce their risk of chronic disease. Users can determine what and how much to eat; track foods, physical activities, and weight; and personalize the tool with goal setting, virtual coaching, and journaling.

With science-based technology drawing on years of research, the Body Weight Planner will enable SuperTracker’s more than 5.5 million registered users to tailor their plans to reach a goal weight during a specific time frame, and maintain that weight afterward.
The math model behind the Body Weight Planner, an online tool published by NIH in 2011, was created to accurately forecast how body weight changes when people alter their diet and exercise habits. This capability was validated using data from multiple controlled studies in people.
The planner’s calculations reflect the discovery that the widely accepted paradigm that reducing 3,500 calories will shed one pound of weight does not account for slowing of metabolism as people change their diet and physical activities. More recently, the math model was further validated [http://www.niddk.nih.gov/news/research-updates/Pages/model-counts-calories-tracking-body-weight.aspx] using data from a two-year calorie-restriction study of 140 people. With those data, the researchers showed the model can also provide accurate measurements of calorie-intake changes by tracking people’s weight. Researchers are examining how to apply this method for public use.
NIEHS: RGS2 PROTEIN HELPS PREPARE HEALTHY EGG-SPERM UNION
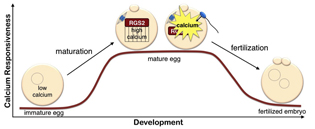
NIEHS researchers, collaborating with colleagues at the University of Connecticut Health Center, the University of Western Ontario, and the University of South Florida, have discovered that the regulator of G-protein signaling–2 (RGS2) protein functions as a brake to suppress premature Ca2+ release in eggs that are poised on the brink of development. While other researchers have shown that the RGS2 protein plays an important role in regulating heart function and blood pressure, this is the first demonstration of the protein’s significant role in fertilization.
During the maturation process, the egg stores calcium, preparing it for fertilization. At fertilization, the sperm causes calcium to be released in the egg, turning it into a developing embryo. The mouse study showed that during maturation, the egg synthesizes RGS2, which suppresses calcium signaling. This safety mechanism ensures that the egg does not begin releasing calcium and start developing before the sperm arrives, because that would prevent the egg from merging with the sperm.
RGS2 is being studied as a therapeutic target for hypertension and other heart ailments. Understanding the role RGS2 plays in reproduction is important when considering the possible benefits and side effects of any new treatments, as well as in understanding the impact that toxins might have on human fertility. (NIH authors: M.L. Bernhardt, E. Padilla-Banks, C.E. McDonough, and C.J. Williams, Development 142:2633–40, 2015)
This page was last updated on Monday, April 25, 2022
