Putting the "I" in RNAi
A Powerful Tool Lets Researchers Get Personal
NICHD researcher Mel DePamphilis hates cancer. One year ago, his then 39-year-old daughter, Kimberly—married with two children and a loving husband—was diagnosed with breast cancer and underwent a double mastectomy. In the months that followed, she received intense chemotherapy and radiation treatments in the hope of destroying the last vestige of her disease. “Cancer touches so many lives,” he rasped, overcome with emotion as he related the story. He took a breath and steadied himself, his eyes narrowed in determination. For DePamphilis, the battle against cancer was no longer academic; it was personal.
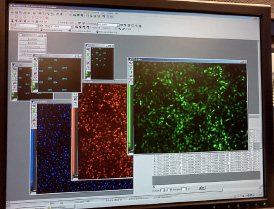
Photo: Nicholas DiCrosta

Photo: Carleen Klumpp, NHGRI
NIH’s RNAi Screening Facility uses a powerful tool called RNA interference (RNAi) that systematically inactivates genes so scientists can probe their functions. The images (top) such as this high-content analysis of a fluorescent virus are produced by the automated RNAi screening system (bottom).
DePamphilis is well positioned to fight that battle. Based on a competitive proposal, he was given access to a sophisticated resource at the NIH Chemical Genomics Center’s (NCGC) RNAi Screening Facility, in Rockville, Md. RNA interference (RNAi), a powerful tool that systematically inactivates genes, enables scientists to determine the functions of genes. The tool is useful for research in many areas including cancer, age-related disorders, metabolic diseases, and disease-causing viruses.
Using RNAi, DePamphilis determined that when geminin—a nuclear protein that is crucial for cancer cells to duplicate their genome—is suppressed, cancer cells commit apoptosis, or cellular suicide. The proliferation of normal cells, however, is not affected. DePamphilis’s work is among the first to demonstrate the power of NIH’s RNAi facility and its potential to help scientists in their quest to defeat disease.
The seeds for the RNAi screening facility were planted in the fall of 2007 when the institutes and centers’ (ICs) scientific directors convened a committee to study how RNAi research could be advanced at NIH. The committee, made up of representatives from several ICs and chaired by Brian Oliver (NIDDK), recommended that the NIH intramural research program construct its own RNAi screening facility. “We knew we had to think big to achieve big things,” Oliver said. “The technology was advancing so rapidly [that] if we didn’t make a move quickly we risked losing relevance.”
The RNAi facility was developed through a collaborative, trans-NIH effort by teams of geneticists, biologists, chemists, engineers, and bioinformatics specialists. The facility was located within the NCGC to leverage the NCGC’s existing small-molecule high-throughput screening, informatics, and chemistry infrastructure. RNAi gurus Natasha Caplen (NCI) and Scott Martin (NHGRI) helped get the facility running; Caplen chaired the oversight committee and Martin developed the screening and informatics platform. The state-of-the art facility opened in 2010 and has been performing screens for DePamphilis and other scientists ever since.
Previously, gene-silencing research at the NIH had been limited to labs that were studying individual genes. With some 23,000 genes in the human genome to dig through, the process of shutting them off one at a time to assess their functionality could take decades and cost billions of dollars. The RNAi facility, however, provides intramural researchers with the tools to screen whole genomes and pathways on an industrial scale. RNAi screens often identify multiple genes that control critical processes and pathways in a particular biology being studied.
By using an assay developed in collaboration with Martin, for example, DePamphilis was able to screen 21,584 genes in just four months.
“It gives us true functional data,” said Caplen, whose seminal work in 2001 showed that gene suppression was possible in mammalian cells. Now “I can get information in a week that would have taken years.”
Martin is also enthusiastic about the future of RNAi research. “Do I get excited when I read the results of a screen? You bet I do!” He is thrilled that his “work can one day help a kid suffering from cancer.”
DePamphilis speculates that he may have discovered cancer’s “Achilles’ heel”: Suppressing geminin should be able to kill many forms of cancer. In preliminary tests, he has been able to destroy the same kind of breast cancer cells that affect his daughter. Although after one year of treatment, his daughter is winning her battle against the disease, DePamphilis isn’t about to give up his personal crusade against cancer. He knows his work is just the opening salvo in the fight. He is optimistic that his collaboration with the RNAi facility will lead him and other researchers down the path to defeating his nemesis: “When it comes to cancer, I don’t take prisoners.”
This is just one scientist’s story about using the RNAi Screening Facility to enhance his research. For more information about the facility and how to apply for project support, visit http://rnai.nih.gov and go to the RNAi symposium, October 24, Kirschstein Auditorium in Natcher Building 45, 2:00–4:00 p.m., at the NIH Research Festival (October 24–28; see Announcements). For information on the NIH Chemical Genomics Center, visit http://www.ncgc.nih.gov.
This page was last updated on Monday, May 2, 2022
