Research Briefs
NHLBI, NIAID: HIGHLY INFECTIOUS MEMBRANE-CLOAKED “VIRUS CLUSTERS” TRANSMIT VIRUSES AMONG HUMANS
CREDIT: MEDICAL ARTS, NIH
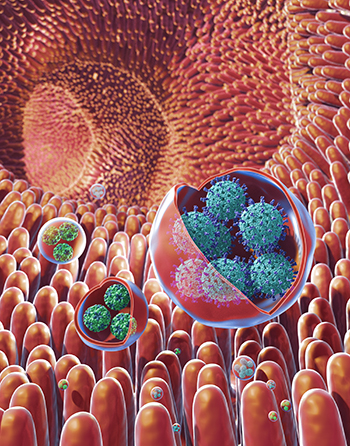
Viruses that cause severe stomach illness are transmitted through membrane-cloaked virus clusters rather than through individual virus particles, according to a new NIH study. Shown: illustration of membrane-bound vesicles containing clusters of viruses, including rotavirus (in the large vesicles) and norovirus (in the small vesicles), within the gut.
NIH researchers have found viruses that cause severe stomach illness—including the one infamous for widespread outbreaks on cruise ships—get transmitted among humans through membrane-cloaked “virus clusters” that exacerbate the spread and severity of disease. Previously, it was believed that these viruses only spread through individual virus particles. The discovery of these clusters, the scientists said, marks a turning point in the understanding of how these viruses spread and why they are so infectious. This preliminary work could lead to the development of more effective antiviral agents than existing treatments that mainly target individual particles.
The researchers studied norovirus and rotavirus, hard-to-treat viruses that are the most common causes of gastroenteritis and that afflict millions of people each year. The viruses cause symptoms ranging from diarrhea to abdominal pain and can sometimes result in death, particularly among young children and the elderly. Their highly contagious nature has led to serious outbreaks in crowded spaces, most notably in cruise ships, daycare centers, classrooms, and nursing homes. Fortunately, vaccines against rotavirus are now available and are routinely given to babies in the United States. Rotaviruses and noroviruses are mainly spread by accidentally ingesting tiny particles of an infected person’s stool—through, for example, contaminated food or liquids.
The researchers obtained fecal samples of humans and animals (pigs and mice) and found that the viruses are shed in the stool as virus clusters inside membrane-bound packets. In addition, they found that these virus-containing vesicles were significantly more infectious than the free, unbound viruses within the samples. The researchers determined that the high level of infectiousness was likely due to the vesicles delivering many viruses at once to the target tissues; protecting their viral cargo from being destroyed by prolonged exposure to enzymes; and possibly by making their viral cargo invisible to the antibodies that are in the stool or gut of the host. More studies are needed, but the extreme potency of the virus packets, they said, has a clear consequence: It not only enhances the virus’ ability to spread more aggressively but it also increases the severity of the disease it causes. Handwashing with soap and water helps prevent the spread of viruses. (NIH authors: M. Santiana, S. Ghosh, B.A. Ho, V. Rajasekaran, W.-L. Du, Y. Mutsafi, D.A. De Jésus-Diaz, S.V. Sosnovtsev, E.A. Levenson, C. Bleck, K.Y. Green, and N. Altan-Bonnet, Cell Host Microbe 24:208–220E8, 2018; DOI:10.1016/j.chom.2018.07.006)
NIAID (HAMILTON, MONTANA): HOW MERS CORONAVIRUS EVOLVES TO INFECT DIFFERENT SPECIES
CREDIT: MICHAEL LETKO, LABORATORY OF VIROLOGY, NIAID
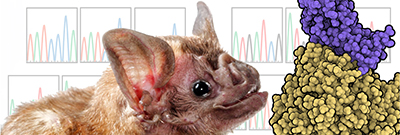
Coronaviruses, which cause severe respiratory disease, originate in bats and evolve so they can be transmitted to other animals and then to humans. NIH scientists found they could observe how the virus evolved by growing it on cells from vampire bats (Desmodus rotundus) that it didn’t normally infect. Shown: The vampire bat species used in the study and an illustration representing Middle East respiratory syndrome coronavirus (purple) interacting with host receptor DPP4 (gold).
In the past 15 years, two outbreaks of severe respiratory disease have been caused by coronaviruses that originated in bats and were passed to humans through other animals. Scientists don’t yet understand the genetic mechanisms underlying cross-species adaptation. New NIAID research shows how Middle East respiratory syndrome coronavirus (MERS-CoV) can adapt to infect cells of a new species, which suggests that other coronaviruses might be able to do the same.
In 2003, severe acute respiratory syndrome coronavirus (SARS-CoV) spread from civets to infect more than 8,000 people, leading to a yearlong global public-health emergency.
MERS-CoV, first identified in 2012, consistently jumps from dromedary camels (Camelus dromedarius) to people, resulting in periodic outbreaks with a roughly 35 percent fatality rate.
While many other coronaviruses are not known to infect people, MERS-CoV and SARS-CoV are notable for their ability to infect a variety of different species, including humans.
To evaluate how MERS-CoV evolves to infect host cells, the scientists tested 16 bat species and found that the virus could not efficiently enter cells with receptors from the common vampire bat, Desmodus rotundus. The researchers then grew virus on cells that had vampire bat receptors and observed the virus evolving to better infect the cells. After a few generations, the virus had completely adapted to the vampire bat receptor. By studying how the shape of MERS-CoV changed over time to attach to the new host receptor, the scientists found similarities with prior studies of SARS-CoV. Thus, while these two viruses are different, they use the same general approach to enter the cells of new species.
Understanding how viruses evolve to infect new species will help scientists determine what is required for viruses to emerge and spread in new hosts and may be important for developing new vaccines, which viruses often evolve to avoid. The NIAID scientists next plan to work with other, related viruses to determine whether they also efficiently adapt to new species. (NIH authors: M. Letko, K. Miazgowicz, R. McMinn, S.N. Seifert, A. Carmody, N. van Doremalen, and V. Munster, Cell Rep 24:1730–1737, 2018; DOI:10.1016/j.celrep.2018.07.045)
NIAMS AND NIDCR: WHY WOUNDS HEAL QUICKER IN THE MOUTH THAN ON THE SKIN
Oral wounds heal more rapidly and with little scarring than wounds on the skin. But the intrinsic characteristics that mediate optimal healing at mucosal surfaces are poorly understood, particularly in humans. Researchers from NIAMS and NIDCR identified the physiological and molecular determinants for this repair paradigm. They took biopsies from the cheeks and arms of 30 people and then took biopsies again two or five days later. Using molecular profiling, the scientists determined that wound-activated transcriptional networks are present at the basal state in the oral mucosa, priming the epithelium for wound repair. Several genes appear to play a key role in the wound healing in the mouth including the SOX2 and PITX1, genes for transcriptional regulators. The researchers found that activating the regulators in mouse skin cells improved skin healing. The findings could have widespread implications for the wound-healing field. (NIH authors: R. Iglesias-Bartolome, A. Uchiyama, A.A. Molinolo, L. Abusleme, S.R. Brooks, J.L. Callejas-Valera, D. Edwards, C. Doci, N.M. Moutsopoulos, J.S. Gutkind, M.I. Morasso, Sci Transl Med 10:eaap8798; DOI:10.1126/scitranslmed.aap8798)
NIDDK, CC: A DRUG TO TREAT OVERACTIVE BLADDER HAS OTHER METABOLIC EFFECTS
An NIDDK study showed that a class of medications, beta-3 adrenergic receptor (ADRB3) agonists, may have multiple physiological effects on human metabolic functioning. The researchers found that the ADRB3 agonist mirabegron, an FDA-approved drug to treat only overactive bladder, increased brown-fat metabolic activity, white-fat lipolysis, and resting-energy expenditure in 12 healthy men. (Only men were recruited for this study because the 200 mg dose used might have caused heart rhythm problems in women; another study is underway to test a 100 mg dose in women.) Mirabegron also affected gallbladder size and bile-acid metabolism, two functions previously unreported in studies of the drug in either rodents or humans. All these effects were substantial only at a dosage higher than that approved to treat overactive bladder, which may explain why many of the physiological responses to ADRB3 agonists in humans have not yet been reported.
Clinical studies have for decades used ADRB3 agonists to treat obesity, yet no ADRB3 agonist has yet been approved for this purpose. The current study highlights the need for further research on ADRB3 agonists’ physiological effects in humans, with the goal of developing more-selective human ADRB3 agonists to treat obesity and other metabolic conditions. The discovery of mirabegron’s effects on gallbladder size and bile acids may also inform future research on ADRB3 agonists’ potential benefits to gastroenterological functioning and the gut microbiome. (NIH authors: A.S. Baskin, J.D. Linderman, R.J. Brychta, S. McGehee, E. Anflick-Chames, C. Cero, J.W. Johnson, A.E. O’Mara, L.A. Fletcher, B.P. Leitner, C.J. Duckworth, S. Huang, H. Cai, H.M. Garraffo, C.M. Millo, W. Dieckmann, P.J. Walter, P. Herscovitch, K.Y. Chen, and A.M. Cypess, Diabetes Jul;db180462, 2018; DOI:10.2337/db18-0462)
NCI: PREDICTOR FOR IMMUNOTHERAPY RESPONSE IN MELANOMA
In a new study, NCI researchers and colleagues at three universities developed a gene-expression predictor that can indicate whether melanoma in a specific patient is likely to respond to treatment with immune checkpoint inhibitors, a novel type of immunotherapy.
Treatment with checkpoint inhibitors is effective for some patients with late-stage melanoma and certain other types of cancer, but not all patients with melanoma respond. In this study, the investigators developed a predictor by first looking for clues in cases in which the immune system appears to mount an unprompted, successful immune response to cancer, causing spontaneous tumor regression. They analyzed neuroblastoma, a type of cancer that frequently undergoes spontaneous regression in young children, and were able to define gene-expression features that separated patients with nonregressing disease from those with regressing disease. The researchers computed an immuno-predictive score (IMPRES) for each patient sample. The higher the IMPRES score for a sample, the more likely the sample was to undergo spontaneous regression. To see whether IMPRES could be used to predict melanoma patients’ responses to checkpoint inhibitors, the authors analyzed 297 samples from several studies. They found that the predictor could identify nearly all patients who responded to the inhibitors and more than half of those who did not, making it significantly superior to all other existing published predictors. Importantly, unlike other existing predictors, IMPRES was accurate across many different melanoma patient data sets.
According to the researchers, the results need to be carefully evaluated in additional patient datasets; further study is warranted in other cancer types for which checkpoint inhibitors have been approved. (NIH authors: N. Auslander, J.S. Lee, S. Madan, and E. Ruppin, Nat Med; DOI:10.1038/s41591-018-0157-9)
NIDCD: THAT STINKS! ONE IN 15 AMERICANS SMELL ODORS THAT AREN’T THERE
CREDIT: EHSTOCK, THINKSTOCK.COM
One in 15 Americans (or 6.5 percent) over the age of 40 experiences phantom odors, according to a new study led by researchers from NIDCD. Problems with the sense of smell can have an impact on appetite, food preferences, and the ability to smell danger signals such as fire, gas leaks, and spoiled food.
The team used data from 7,417 participants over 40 years of age from CDC’s 2011–2014 National Health and Nutrition Examination Survey (NHANES).
Researchers used this NHANES survey question: “Do you sometimes smell an unpleasant, bad, or burning odor when nothing is there?” To explore the correlation between phantom odors and participant characteristics, the researchers looked at participants’ age, sex, education level, race/ethnicity, socioeconomic status, certain health habits, and general health status. The ability to identify odors in the environment is known to decrease with age. This study found that the prevalence of phantom odor perception also decreases with age and is not related to individuals’ ability to correctly identify odors. The study also found that about twice as many women as men reported phantom odors, and that the female predominance was particularly striking for those under age 60.
Other risk factors for the onset of phantom odors include head injury, dry mouth, poor overall health, and low socioeconomic status. Researchers hypothesized that people with lower socioeconomic status may more commonly be exposed to environmental pollutants and toxins or have health conditions that contribute to phantom odors, either directly or because of medications needed to treat their health conditions. The study could inform future research aiming to unlock the mysteries of phantom odors. (NIH author: K.E. Bainbridge, JAMA Otolaryngol Head Neck Surg; DOI:10.1001/jamaoto.2018.1446)
NICHD, NCI: BLOOD TEST MAY IDENTIFY GESTATIONAL DIABETES RISK IN FIRST TRIMESTER
Pregnant women are typically screened for gestational diabetes between 24 and 28 weeks of pregnancy, but a new study by NIH researchers suggests that a blood test at 10 weeks may diagnose the condition sooner. The researchers evaluated whether the hemoglobin A1c (HbA1c) test (also called the A1C test), commonly used to diagnose type 2 diabetes, could identify signs of gestational diabetes in the first trimester of pregnancy. The test is not currently recommended to diagnose gestational diabetes at any point in pregnancy. The study involved analyzing records from the NICHD Fetal Growth Study, a large observational study that recruited more than 2,000 low-risk pregnant women from 12 U.S. clinical sites between 2009 and 2013. The researchers compared HbA1c test results from 107 women who later developed gestational diabetes with test results from 214 women who did not develop the condition. Women who went on to develop gestational diabetes had higher HbA1c measurements (an average of 5.3 percent) than those without gestational diabetes (an average HbA1c of 5.1 percent). Each 0.1 percent increase in HbA1c above 5.1 percent in early pregnancy was associated with a 22 percent higher risk for gestational diabetes.
Further studies are needed to confirm the findings and to determine whether lowering HbA1c with lifestyle changes—such as exercise and healthy diet—either in early pregnancy or before pregnancy, could reduce the risk for the condition. (NIH authors: S.N. Hinkle, S. Rawal, P.S. Albert, and C. Zhang, Sci Rep 8:12249, 2018; DOI:10.1038/s41598-018-30833-8)
NIDCD: NOVEL DRUG THERAPY PARTIALLY RESTORES HEARING IN MICE
CREDIT: YOKO NAKANO, CARVER COLLEGE OF MEDICINE, UNIVERSITY OF IOWA
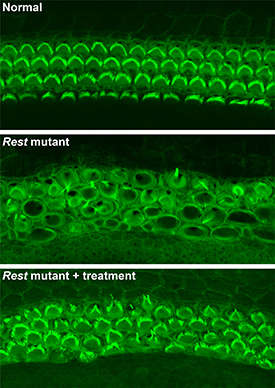
NIH and University of Iowa researchers used a small-molecule drug to partially restore hearing in deaf house mice. Shown: (top) Rows of healthy sensory hair cells with green stereocilia arcs in the mouse inner ear; (middle) In Rest-mutant mice, hair cells are disorganized, and stereocilia are barely visible; (bottom) In Rest-mutant mice treated with the drug, the organization and structure of hair cells is partially restored.
A small-molecule drug is one of the first to preserve hearing in a house mouse (Mus musculus) model of an inherited form of progressive human deafness, reported investigators from NIDCD and the University of Iowa (Iowa City, Iowa). The study sheds light on the molecular mechanism that underlies an inherited form of deafness called DFNA27 and suggests a new treatment strategy.
The seed for the advance was planted a decade ago when NIDCD researchers (two of whom were authors on the current study) localized the deafness-causing mutation in a region on chromosome four called DFNA27, which includes a dozen or so genes. The precise location of the mutation eluded the NIDCD team, however, until now. A crucial clue to explain the DFNA27 form of progressive deafness arose from studies of the mouse Rest gene, which codes for the protein REST (neuron-restrictive silencer element–silencing transcription factor), conducted by researchers at the University of Iowa. When the University of Iowa team deleted exon 4 of Rest in mice, inner-ear hair cells died, and mice became deaf. Many genes that should have been active were shut off in hair cells before their death. Working together, the NIH researchers and the Iowa group pinpointed the deafness mutation and determined that it lies near exon 4, alters the boundaries of exon 4, and interferes with the inactivation of REST in hair cells. Using small-molecule drugs that inhibit this process, the investigators were able to turn off REST and partially restore hearing. According to the researchers, if additional studies show that small-molecule-based drugs are effective in treating DFNA27 deafness in people, it’s possible that using similar approaches might work for other inherited forms of progressive hearing loss, too. (NIH authors: M.C. Kelly, A.U. Rehman, E.T. Boger, R.J. Morell, M.W. Kelley, and T.B. Friedman, Cell 174:536–548, 2018; DOI:https://doi.org/10.1016/j.cell.2018.06.004)
NICHD: HOW MALARIA PARASITES ARE RELEASED FROM RED BLOOD CELLS
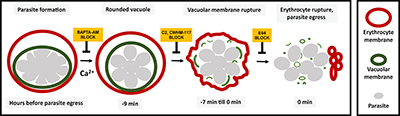
CREDIT: SVETLANA GLUSHAKOVA, NICHD
Diagram showing the sequence of events involved in rupture of the vacuole and host cell membrane leading to release of the malaria parasite. Using chemical inhibitors, the researchers showed that it’s possible to block each event in the sequence.
The vacuole, a compartment inside human red blood cells in which malaria parasites reproduce and develop, takes on a distinct spherical shape just minutes before its membrane ruptures, leading to the release of parasites into the blood stream, according to researchers at NICHD and other institutions. Working with red blood cells from healthy donors, the researchers were able to chemically block the sequence of events that leads to this rounding of the vacuole. They noted that targeting this sequence could inform new treatment strategies against Plasmodium falciparum, the species of malaria parasite that causes the most deaths worldwide and, in several areas, has become drug-resistant.
To track the rounding sequence under a microscope, researchers dyed the membrane of the vacuole with a substance that gives off green light. About 10 minutes before the membrane ruptured, the vacuole morphed from a lumpy, uneven shape to a sphere. Previous studies showed that malaria parasites use calcium to trigger the biochemical reactions needed for their release from the cell. When the researchers treated the cells with a compound that blocks calcium’s effect, the vacuoles couldn’t transition to the spherical form, trapping the parasites inside the cell. (NIH authors: S. Glushakova, M. Garten, B.L. Busse, T. Tenkova‐Heuser, J. Heuser, and J. Zimmerberg, Cell Microbiol e12868, 2018; DOI:10.1111/cmi.12868)
NCI: LOW-DOSE RADIATION EXPOSURE LINKED TO LEUKEMIA
Using data from nine historical cohort studies, investigators in NCI’s Radiation Epidemiology Branch and colleagues from other institutions were able to quantify—for the first time—excess risk for leukemia and other myeloid malignancies that occurred after low-dose exposure to ionizing radiation in childhood. Given the ubiquity of exposure, primarily from medical procedures such as computed tomography scans, every effort should be made to minimize doses, especially for children, according to the researchers. Data for this analysis came from more than 250,000 people aged 21 or younger at the time of first exposure and were contributed from nine cohort studies (from Canada, France, Japan, Sweden, the United Kingdom, and the United States) enrolled between June 4, 1915, and December 31, 2004. Evidence from this study suggests that there is significant risk even at these lower doses and that the current system of radiological protection is prudent and not overly protective. (NIH authors: M.P. Little, D. Borrego, C. Lee, A.V. Brenner, D. Campbell, M.M. Doody, M.S. Linet, and A. Berrington de González, Lancet Haematol 8:e346-e358, 2018; DOI:10.1016/S2352-3026(18)30092-9)
NIAID: SCIENTISTS CREATE 3-D STRUCTURE OF 1918 INFLUENZA VIRUS-LIKE PARTICLES
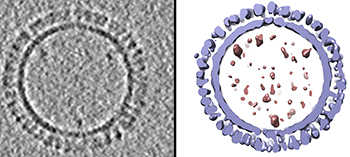
CREDIT: DUSTIN MCCRAW AND AUDRAY HARRIS, NIAID
On the left is a 1918 H1 influenza virus-like particle (VLP) as seen by cryoelectron microscopy. On the right is the same VLP rendered in 3-D with structural components computationally segmented and colored; hemagglutinin and membrane are light blue and internal components (molecular cargo) are red.
A team of NIAID researchers who specialize in visualizing molecular structures developed a 3-D model of virus-like particles (VLPs) based on the 1918 H1 pandemic influenza virus. VLPs are noninfectious protein-based structures that bind to antibodies and show promise as vaccine platforms for many viral diseases including influenza.
Other researchers had produced VLPs for 1918 H1 influenza that successfully protected animals from different influenza viruses. The NIAID group prepared hundreds of such VLP samples and analyzed their structure with cryoelectron microscopy, a technique that quick-freezes samples with glasslike clarity. The researchers then sliced through those VLP 3-D structures to analyze their internal structure, using computers to document the size and placement of key molecules. After averaging all their data, the group then created a 3-D 1918 influenza VLP model.
It turned out that about 90 percent of the VLPs are hemagglutinin (HA) proteins (by weight) found on the VLP surface. In contrast, HAs are less than half of the viral proteins of natural influenza viruses. The number and location of HA molecules may influence the efficacy of VLP vaccines, influencing the binding of antibodies to specific epitopes on the HA protein. Those antibodies can similarly bind live influenza viruses, preventing them from infecting cells. The more that is understood about the molecular organization of influenza VLPs, the better scientists will be able to develop effective seasonal and universal influenza vaccines. (NIH authors: D.M. McCraw, J.R. Gallagher, U. Torian, M.L. Myers, M.T. Conlon, N.M. Gulati, and A.K. Harris, Sci Rep 8:10342, 2018; DOI:10.1038/s41598-018-28700-7)
NICHD: INDUCED LABOR AT 39 WEEKS MAY REDUCE LIKELIHOOD OF C-SECTION
Healthy first-time mothers whose labor was induced in the 39th week of pregnancy were less likely to deliver by cesarean section (C-section) compared with those who waited for labor to begin naturally, according NICHD researchers. The researchers also found that infants born to women induced at 39 weeks were no more likely to experience stillbirth, newborn death, or other severe complications than infants born to uninduced women. The study enrolled more than 6,000 pregnant women at 41 hospitals participating in the NICHD-supported Maternal-Fetal Medicine Units Network. Roughly half of the women were assigned at random to have their labor induced in the 39th week of pregnancy; the remaining women received expectant management. The researchers estimate that one cesarean delivery could be avoided for every 28 low-risk, first-time mothers undergoing elective induction at 39 weeks. (NIH authors: W.A. Grobman, M.M. Rice, U.M. Reddy, A.T.N. Tita, R.M. Silver, G. Mallett, K. Hill, E.A. Thom, Y.Y. El-Sayed, A. Perez-Delboy, Dw.J. Rouse, G.R. Saade, K.A. Boggess, S.P. Chauhan, J.D. Iams, E.K. Chien, B.M. Casey, R.S. Gibbs, S.K. Srinivas, G.K. Swamy, H.N. Simhan, and G.A. Macones, N Engl J Med 379:513–523, 2018; DOI:10.1056/NEJMoa1800566)
NIAID, CC: IMAGING TECHNIQUE ILLUMINATES IMMUNE STATUS OF MONKEYS WITH HIV-LIKE VIRUS
Findings from an animal study suggest that a noninvasive imaging technique could, with further development, become a useful tool to assess immune-system recovery in people receiving treatment for HIV infection, according to new research conducted by NIAID and CC scientists. The researchers used single-photon emission computed tomography and a cluster of differentiation–4 (CD4)–specific imaging probe to assess immune system changes—pools of CD4 T cells in tissues such as lymph nodes, spleen, and gut—throughout the bodies of rhesus macaques (Macaca mulatta) infected with SIV, a simian form of HIV, after the initiation and interruption of antiretroviral therapy (ART). The findings illustrate that CD4+ T-cell concentrations in the blood—a measure of immune-system health in people living with HIV—often fail to fully reflect the situation in tissues. A low blood CD4+ T-cell concentration indicates an immune system weakened by HIV, and the level generally increases when ART is started and the immune system begins to recover. In the future, extension of this imaging technology to humans may aid scientists in evaluating immune reconstitution after standard and experimental HIV treatments, the authors conclude. (NIH authors: M. Di Mascio, S. Srinivasula, I. Kim, G. Duralde, A. St. Claire, P. DeGrange, M. St. Claire, E.E. Gabriel, R.C. Reba, C. Paik, and H.C. Lane, JCI Insight 3:e97880, 2018; DOI:10.1172/jci.insight.97880)
NICHD: TEEN CRASH RISK HIGHEST DURING FIRST THREE MONTHS AFTER GETTING DRIVER’S LICENSE
Teenage drivers are eight times as likely to be involved in a collision or a near miss during the first three months after getting a driver’s license as during the previous three months on a learner’s permit, according to a study led by NICHD researchers. Teens are also four times as likely to engage in risky behaviors, such as rapid acceleration, sudden braking, and hard turns, during this period. In contrast, teens on a learner’s permit drove more safely, with their crash or near-crash and risky driving rates similar to those of adults. The study enrolled 90 teenagers and 131 parents in Virginia, and the data-collection system was developed by the Virginia Tech Transportation Institute (Blacksburg, Virginia). The study is one of the first to follow the same individuals over time, from the beginning of the learner period through the first year of independent driving, while continuously collecting information using software and cameras installed in the participants’ vehicles. The findings suggest that teen drivers may benefit from a more gradual decrease in adult supervision during the first few months of driving alone. (NIH authors: P. Gershon, C. Zhu, K.R. Sita, and B. Simons-Morton, J Adolesc Health 18:30189–30187; DOI:10.1016/j.jadohealth.2018.04.012)
NIAID (BETHESDA AND HAMILTON, MONTANA) NATURAL LIPID ACTS AS POTENT ANTI-INFLAMMATORY
NIAID researchers have identified a naturally occurring lipid used by a disease-causing bacterium to impair the host immune response and increase the chance of infection. Inadvertently, they also may have found a potent inflammation therapy against bacterial and viral diseases. Lipids are known to help Francisella tularensis bacteria, the cause of tularemia, to suppress host inflammation when infecting mouse and human cells. In the new study, NIAID researchers found a form of the lipid phosphatidylethanoloamine (PE) present in the bacterium. The composition of PE found in F. tularensis differs from PE found in other bacteria. In cell-culture experiments, the researchers discovered that the natural and a synthetic form of PE reduced inflammation caused by both tularemia bacteria and dengue fever virus. The group hopes their findings will eventually lead to the development of a potent, broad-spectrum anti-inflammatory therapeutic. (NIH authors: R. Ireland, B. Schwarz, G. Nardone, T.D. Wehrly, A.I. Chiramel, S.M. Best, and C.M. Bosio, J Innate Immun; DOI: 10.1159/000489504)
NIAID: MECHANISMS LEADING TO DIFFERENT B-CELL FATES HAVE BEEN UNCOVERED
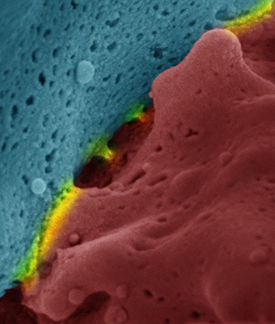
CREDIT: BILLUR AKKAYA, NIAID
Scanning electron microscope image of an immune synapse. Red and blue show the interacting cells and yellow highlights the surfaces of interaction.
NIAID researchers have discovered novel mechanisms that dictate the survival, death, or differentiation of the antibody-producing B cells that have encountered a foreign antigen. The scientists showed that signaling cascades initiated by the binding of the antigen to the B-cell receptor (BCR) lead to rapid increases in cellular respiration, nutrient uptake, and expression of various activation markers. These early changes prepare the B cell to further differentiate into antibody-producing cells. However, unless a second, independent stimulus is received soon after antigen binding, BCR signaling does not lead to B-cell proliferation and differentiation. Instead, in the absence of a second signal, antigen binding to the BCR leads to a dysregulation of cellular calcium homeostasis that in turn leads to mitochondrial dysfunction and a phenomenon called activation-induced cell death. The second signal can be provided by toll-like receptor (TLR) signaling to pathogen products, indicating the presence of an acute infection, or through B cell–T cell interactions that rule out that the initial antigen was a prohibited self-antigen that would lead to autoimmunity. (NIH authors: M. Akkaya, J. Traba, A.S. Roesler, P. Miozzo, B. Akkaya, B.P. Theall, H. Sohn, M. Pena, M. Smelkinson, J. Kabat, E. Dahlstrom, D.W. Dorward, J. Skinner, M.N. Sack, and S.K. Pierce, Nat Immunol 19:871–884, 2018; DOI:10.1038/s41590-018-0156-5)
In another study, the researchers uncovered additional details about the interplay of TLR- versus T-cell-mediated second signals, namely, that TLR signaling trumped the need for the antigen-stimulated B cell to interact with T cells to receive a “go” signal. TLR stimulation decreased the B cells’ ability to capture, process, and present antigens to solicit help from T cells. Rather, TLRs directly boosted B-cell antibody and cytokine secretion and proliferation. Thus, TLR signals drive B cells toward rapid T-cell-independent differentiation to plasma cells, providing rapid neutralizing antibody production to fight off infections quickly. In contrast, B cells that receive their survival signals from T cells are induced to undergo a time-consuming process that is necessary for antibody affinity maturation and long-lasting immune memory. Therefore, the TLR-driven shortcut to quick antibody comes with a price, which is the absence of “crème de la crème” antibodies that would protect the body from subsequent infections for years to come. (NIH authors: M. Akkaya, B. Akkaya, A.S. Kim, P. Miozzo, H. Sohn, M. Pena, A.S. Roesler, B.P. Theall, T. Henke, J. Kabat, J. Lu,D.W. Dorward, E. Dahlstrom, J. Skinner, L.H. Miller, and S.K. Pierce, Nat Immunol 19:255–266, 2018; DOI:10.1038/s41590-018-0052-z)
Commentary on both studies: Nat Immunol 19:791–793, 2018)
NIAID, CC: DURING HIV INFECTION, ANTIBODY CAN BLOCK B CELLS FROM FIGHTING PATHOGENS
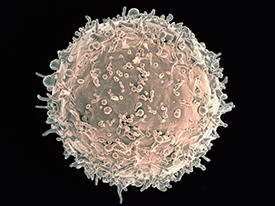
CREDIT: NIAID
Colorized scanning electron micrograph of a B cell from a human donor.
For the first time, scientists have shown that in certain people with HIV, the antibody immunoglobulin G3 (IgG3) stops the immune system’s B cells from fighting pathogens. This phenomenon appears to be one way the body tries to reduce the potentially damaging effects of immune-system hyperactivity caused by the presence of HIV, according to the NIAID and CC investigators who led the research. However, in so doing, the phenomenon also impairs normal immune function.
The scientists made their discovery by analyzing blood samples from 83 anonymous donors who were not infected with HIV and 108 people with HIV at various stages of infection. The people with HIV came from a variety of racial and ethnic backgrounds. Some were being treated for their infection, while others had not yet begun therapy. IgG3 appeared on the surface of B cells only under certain conditions: in people with HIV, but not in HIV-uninfected people; and on B cells of people of African-American or black African descent during the chronic phase of untreated HIV infection when the virus was not adequately controlled. (NIH authors: L. Kardava, H. Sohn, C. Youn, J.W. Austin, W. Wang, C.M. Buckner, J.S. Justement, V.A. Melson, G.E. Roth, M.A. Hand, K.R. Gittens, R.W. Kwan, M.C. Sneller, T.-W. Chun, P.D. Sun, S.K. Pierce, and S. Moir, Nat Immunol 19:1001–1012, 2018; DOI:10.1038/s41590-018-0180-5)
This page was last updated on Wednesday, April 6, 2022
