Scientists with TALEN(t)!
By manipulating the mouse genome, Rafael Casellas brings human gene therapy closer to reality.

Research fellow Kyongrim Kwon, Ph.D., and Principal Investigator Rafael Casellas, Ph.D., create tools for genetic modification
Treatment for X-linked severe combined immunodeficiency (SCID), also known as “bubble boy” disease, uses viruses to insert intact copies of the IL2RG gene into defective cells containing a defective genetic mutation. But for some children with X-linked SCID, this strategy resolves their life-threatening disease while also causing leukemia.

Research assistant Gustavo Gutierrez, Ph.D., and visiting fellow Arito Yamane, M.D., Ph.D., use high-throughput sequencing to gather data from experiments at the bench
With more than 4,000 known single-gene disorders and only a handful of therapies available to treat them, hopes for effective gene therapy continue to hover on the horizon—and that’s the problem: gene therapy has been on the horizon for a long time. Why aren’t we there yet?
“The issue,” says Rafael Casellas, Ph.D., “is that, despite major advances in our understanding of DNA manipulation, gene therapy still relies on random integration of the therapeutic gene.”
Since the discovery of recombinant DNA techniques in the 1970s, DNA manipulation has grown substantially more sophisticated, yet the ability to consistently modify the genome at will continues to elude researchers. Therapeutic viruses typically integrate very well into host DNA, but they do so randomly, which can result in unintended consequences.
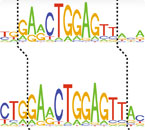
A snippet from a genetic sequence, with the height of each letter corresponding to how frequently the nucleotide is present in the genome
The goal is to clearly define a gene or area of the genome for alteration in a specific way and then target that gene with absolute specificity, either by therapeutically “repairing” a single genetic mutation or by inducing a single mutation for research purposes.

Staff scientist Steven Fellini, Ph.D., manages the Biowulf cluster, a supercomputer used to process massive amounts of experimental genomic data
Tackling the basic problem of specificity in their studies of the mouse genome, Casellas and his team experiment with sequence-specific nucleases and other novel recombination tools.
Like a string of letters in a word-processing program, DNA can be cut, copied, and pasted. Nucleases are biological “scissors” used to cut double-stranded DNA and expose its two phosphate backbones, which can then be modified and rejoined.
When double-stranded DNA is cut, there are two main biological mechanisms that connect the pieces: homologous recombination (HR) and non-homologous end joining (NHEJ). HR is a near-perfect repair mechanism that usually restores function. On the other hand, NHEJ is more common than HR, but is also prone to error and often introduces genetic mutations.

Staff scientist Wolfgang Resch, Ph.D., analyzes data from the lab for information that will lead to new insights about the genome
“Specificity is essential,” Casellas says. “Without specificity, nucleases generate thousands of random strand breaks, and although you may get a break in the right place in one or two cells, that’s not going to be sufficient for therapeutic use or experimental analysis.”

Building specialized Transcription Activator-Like Effector Nucleases (TALENs) by hand
Casellas and colleagues have turned to Transcription Activator-Like Effector Nucleases (TALENs) and a new RNA-guided system called CRISPR in their search for specificity. These synthetic nucleases, coupled to proteins that are engineered to recognize very specific sequences of DNA, can accurately cleave DNA molecules to researchers’ specifications.
Creating each pair of molecular scissors takes about one week in the lab, so Casellas enlists robots to help. When supplied with a large set of defined DNA pieces (plasmids), a robot can efficiently assemble hundreds of customized molecular scissors in a month.

Staff scientist Stevenson Nelson, Ph.D., uses a robot to speed up the assembly of heavily used TALENs
The team now investigates how best to ensure that the repair, or “paste,” element of the modification is done via the high-fidelity HR method and not with error-prone NHEJ. There are ways of tricking cells to do this, but Casellas wants longer-term solutions that could set the standard in ensuring accurate replacement of mutated, damaged, or missing genes.
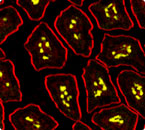
Plasma cells (yellow) from mouse intestinal villi of transgenic mice developed in the Casellas lab
For Casellas, the basic science underpinning their work holds promise in many more ways – accurate targeting of genetic elements is a staple of biomedical research laboratories throughout the world, and as knowledge of the tools expands so too do opportunities for sharing and collaboration.

In the last two years, Casellas has been implementing TALEN-mediated gene targeting to complement the NIH Mouse Regulome Project, the intramural counterpart of the Encyclopedia of DNA Elements (ENCODE) project that seeks to classify and characterize all gene regulatory domains in the mouse genome. By collaborating with extramural and NIH laboratories, Casellas shares and gains intellect and resources that contribute to improving genetic manipulation across research programs and projects.
Rafael Casellas, Ph.D., is a Senior Investigator in the Laboratory of Molecular Immunogenetics of the National Institute of Arthritis and Musculoskeletal and Skin Diseases (NIAMS) and an adjunct investigator at the National Cancer Institute’s Center for Cancer Research (NCI-CCR).
This page was last updated on Wednesday, May 24, 2023