Bookmark this Gene
Karen Adelman investigates how the body’s cells stay on top of their environment.

Karen Adelman, Ph.D., studies how cells respond to environmental stresses
When Karen Adelman, Ph.D., talks about studying transcriptional responses to the environment, few people understand the full scope of her research, so she always has several probing questions ready to help illustrate the field.
“How do cells respond so rapidly to an extracellular signal, but in an absolutely precise manner?” she asks. “How do hormones or growth factors signal to a cell during development to become an organ? In a fully developed organism, how do immune cells react so quickly to pathogens?”

Drosophila, or fruit flies, are a useful model for studying the effects of genetic changes on transcription during development and immune challenge
The awe in Adelman’s voice, fueled by her desire to understand the “black box” that is a cell nucleus, hints at a relentless quest to answer one of biology’s biggest unresolved mysteries: Just how do signals from outside a cell turn certain genes on or off within the nucleus of a cell?
In recent years, science has found that the genetic material within cell nuclei is packaged into nucleosomes—protein units that help structure DNA into specific conformations—resulting in genetic material being either hidden from or exposed to the transcription machinery. The convoluted packaging of DNA is different in each cell and believed to protect genes from aberrantly being turned on. Adelman’s group focuses on the interplay between the packaging of the genome and a cell’s responsiveness to different extracellular signals.

Postdoctoral fellow Daniel Gilchrist, Ph.D., prepares to analyze a microarray covering the entire Drosophila genome to localize chromatin proteins
To map where the transcription machinery is located within a cell, her team used chromatin immunoprecipitation, or ChIP, analysis. Surprisingly, they found that the key RNA polymerase II enzyme that transcribes the information within the DNA genome widely sits on genes, regardless of their activity level. Strikingly, this RNA polymerase is “paused” in action at many genes, essentially holding the chromatin open—and thereby priming the gene for future expression.

Postdoctoral fellow Telmo Henriques, Ph.D., examines the offspring of two Drosophila genotypes, which will be analyzed for development and behavior changes associated with altered transcriptional responses
“Much like we use bookmarks to help navigate a novel,” Adelman explains, “the cell can use paused RNA polymerase II to bookmark specific genes so that they can be readily switched on or turned up when necessary.”
Similar to other paradigm-shifting findings, Adelman’s discovery of genome-wide transcriptional pausing or bookmarking created more questions than answers. One of the first challenges for her team was to determine the importance of pausing. Are RNA polymerase II bookmarks required for cellular function, or are they just handy for transcriptional efficiency? Do they play a more complex role?
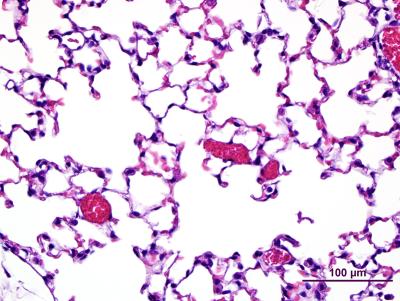
Lung tissue from a normal mouse with open airways (clear spaces) and intact blood vessels (red)
Adelman’s group next dissected the activity of the negative elongation factor (NELF) protein complex, which is necessary for holding RNA polymerase II near the promoter and not letting it escape into the genes. By specifically removing the NELF complex in pluripotent stem cells, the team could impair pausing and ask what the consequences were. Remarkably, they found that NELF-deficient stem cells lose the capacity for self-renewal and do not differentiate.
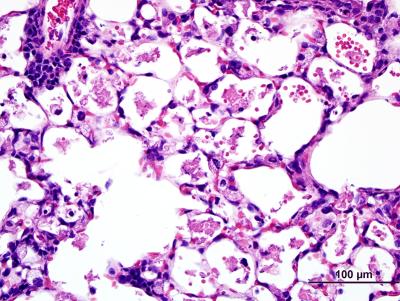
In the absence of immune challenge, lung tissue from mice with macrophages lacking the NELF-B protein reveals signs of inflammation and hemorrhage, suggesting that macrophages lacking NELF activity are hyper-responsive to their environment
“It seems that without pausing, you can’t have life,” Adelman explains.
So what exactly does bookmarking do in cells?
“We’re also using the innate immune system as a model to understand the role of bookmarking by paused RNA polymerase II,” Adelman says. “In this system, we can ask how macrophages switch on protective genes when they are faced with an immune challenge. We already know that certain transcription factors like NF-KB enter into the nucleus and are involved in regulating genes, but we want to understand how they actually flip the switch to turn genes on with different speeds and to different levels.”

Postdoctoral fellow Lucy Williams, Ph.D., uses bone marrow-derived macrophages from mice lacking NELF-B to investigate cytokine production
By removing the pause-inducing factor NELF, she and her team demonstrated a broad mis-regulation of genes involved in regulating the immune pathway, suggesting that the pausing mechanism functions as a rheostat, providing a more delicate level of genetic control than the traditionally suggested “on-off” model. Additionally, finding that reduction of pausing widely affected the responsiveness of immunity genes suggests that modulation of proximal-promoter pausing—or bookmarking—could one day have therapeutic potential.

Dr. Adelman and biologist Ginger Muse working at the bench
Today, Adelman plays a central role in an unfolding scientific story, as research into genetic switches that govern every aspect of human biology continues. After making the initial discoveries in a new and exciting realm of science, does she feel pressure to make breakthrough findings on a daily basis?
“People have been incredibly supportive of my science, knowing that I’m going after some really complex questions,” Adelman says. “Their support gives me the confidence to know I can crack these questions.”
In the meantime, we’ll bookmark this page and prepare for the next chapter in Adelman’s research.
Karen Adelman, Ph.D., is Principal Investigator for the Transcriptional Responses to the Environment Group in the Laboratory of Molecular Carcinogenesis at the National Institute of Environmental Health Sciences (NIEHS).
This page was last updated on Wednesday, May 24, 2023